Oncogenes & TSGs - Accelerators & Brakes
- Oncogenes: "Accelerators" of cell growth.
- Arise from proto-oncogenes via gain-of-function mutations.
- Dominant: one hit sufficient.
- Examples: RAS, MYC, HER2/neu, ABL, BRAF.
- Tumor Suppressor Genes (TSGs): "Brakes" on cell division.
- Inactivated by loss-of-function mutations.
- Recessive: two hits needed (Knudson's hypothesis).
- Roles: DNA repair, cell cycle control, apoptosis.
- Examples: TP53, RB1, BRCA1/2, APC, PTEN.
⭐ TP53 ("guardian of the genome") is the most commonly mutated gene in human cancers.
 and tumor suppressor gene (brake) mechanisms in cancer cells with examples like RAS and TP53 respectively.)
and tumor suppressor gene (brake) mechanisms in cancer cells with examples like RAS and TP53 respectively.)
Hallmarks of Cancer - Rebel Cell Playbook
- Sustaining proliferative signaling: Uncontrolled growth signals (e.g., EGFR, RAS).
- Evading growth suppressors: Ignoring stop signals (e.g., TP53, RB inactivation).
- Resisting cell death: Bypassing apoptosis (e.g., BCL2 upregulation).
- Enabling replicative immortality: Telomerase activation, unlimited divisions.
- Inducing angiogenesis: Stimulating new blood vessel formation (e.g., VEGF).
- Activating invasion & metastasis: Spreading to distant sites (e.g., E-cadherin loss).
- Reprogramming energy metabolism: Preferential aerobic glycolysis.
- Evading immune destruction: Hiding from immune surveillance (e.g., PD-L1).
- Genome instability & mutation: ↑ mutation rates, defective DNA repair.
- Tumor-promoting inflammation: Inflammatory cells fostering tumor growth.
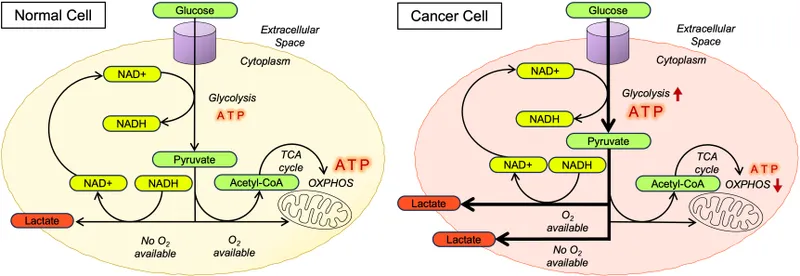
⭐ The Warburg effect describes cancer cells' reliance on glycolysis for energy production, even in the presence of oxygen, providing building blocks for rapid proliferation.
Molecular Diagnostics - Cancer Code Breakers
- Core Techniques & Clinical Utility:
- PCR (Polymerase Chain Reaction): Rapidly amplifies DNA/RNA for mutation detection, MRD.
- Variants: RT-PCR (RNA), qPCR (quantification).
- FISH (Fluorescence In Situ Hybridization): Visualizes gene copy number (e.g., HER2 ↑), translocations (e.g., BCR-ABL).
- IHC (Immunohistochemistry): Detects protein expression guiding therapy (e.g., ER/PR, HER2, PD-L1).
- NGS (Next-Generation Sequencing): Comprehensive genomic analysis; identifies mutations, fusions, CNVs for targeted therapy.
- PCR (Polymerase Chain Reaction): Rapidly amplifies DNA/RNA for mutation detection, MRD.
- Liquid Biopsy: Non-invasive; analyzes ctDNA for diagnosis, monitoring, resistance.

⭐ NGS is pivotal for precision oncology, matching patients with targeted drugs based on tumor molecular profile.
Targeted Therapy - Precision Cancer Strikes
- Drugs designed to interfere with specific molecules ("molecular targets") that drive cancer cell growth and spread.
- Key Principles:
- Identify target (e.g., mutated gene, overexpressed protein).
- Select drug specific to that target.
- Examples:
- Imatinib (BCR-ABL in CML, c-KIT in GIST).
- Trastuzumab (HER2 in breast/gastric cancer).
- Erlotinib/Gefitinib (EGFR mutations in NSCLC).
- Vemurafenib (BRAF V600E in melanoma).
- Requires companion diagnostics for patient selection.
⭐ Resistance to targeted therapies (e.g., T790M in EGFR for NSCLC) is a major challenge, often requiring next-generation inhibitors.
High‑Yield Points - ⚡ Biggest Takeaways
- Oncogenes (e.g., RAS, MYC) promote growth; tumor suppressor genes (e.g., TP53, RB) inhibit it.
- Knudson's two-hit hypothesis explains inactivation of tumor suppressor genes like RB.
- Key Hallmarks of Cancer include sustained proliferation, angiogenesis, and metastasis.
- Liquid biopsies detect circulating tumor DNA (ctDNA) for diagnosis and monitoring.
- Microsatellite instability (MSI) is a feature of Lynch syndrome and some sporadic cancers.
- Epigenetic alterations, like DNA methylation, contribute to carcinogenesis without DNA sequence changes.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

