Basics & Central Dogma - Code of Life
- DNA: Double helix. Bases: Adenine (A), Guanine (G), Cytosine (C), Thymine (T). A=T, G≡C.
- RNA: Single-stranded. Bases: A, G, C, Uracil (U).
- Central Dogma: DNA $\xrightarrow{\text{Replication}}$ DNA $\xrightarrow{\text{Transcription}}$ RNA $\xrightarrow{\text{Translation}}$ Protein.
- Reverse Transcription: RNA → DNA (retroviruses).
- Genetic Code: Triplet codons (3 bases = 1 Amino Acid).
- 64 codons: 61 sense (for AAs), 3 stop (UAA, UAG, UGA). 📌 U Are Away, U Are Gone, U Go Away.
- Start codon: AUG (Methionine).
- Properties: Degenerate, unambiguous, universal (mostly), non-overlapping.
⭐ Degeneracy of the genetic code (multiple codons for one amino acid) acts as a mutation buffer.
Transcription - Scribing the Message
- Process: DNA template → complementary RNA. Enzyme: RNA Polymerase.
- Prokaryotes: Single RNA Pol. Holoenzyme (σ factor for promoter recognition).
- Eukaryotes:
- RNA Pol I: rRNA (28S, 18S, 5.8S). Located in Nucleolus.
- RNA Pol II: mRNA, snRNA, miRNA. Nucleoplasm. (📌 Most Sensitive to α-amanitin)
- RNA Pol III: tRNA, 5S rRNA, U6 snRNA. Nucleoplasm.
- Key Steps:
- Initiation: RNA Pol & General Transcription Factors (GTFs) bind promoter (e.g., TATA box ~-25 bp, CAAT box ~-75 bp). Forms initiation complex.
- Elongation: RNA synthesis in 5'→3' direction. Template strand read 3'→5'.
- Termination: Specific termination sequences signal release of RNA.
- Regulation: Enhancers (↑ rate, can be distant), Silencers (↓ rate).
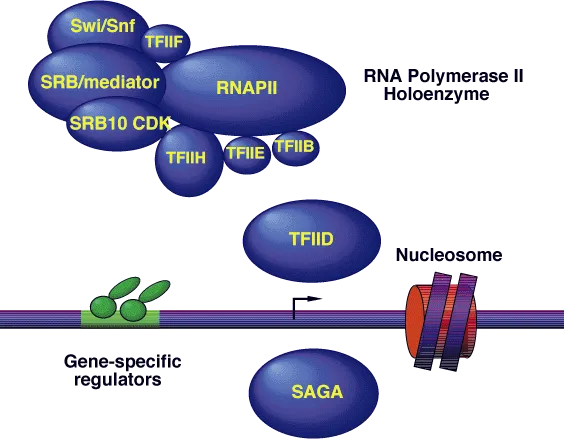
⭐ Rifampicin inhibits prokaryotic DNA-dependent RNA polymerase (β subunit), crucial for TB treatment.
RNA Processing & Translation - Message to Protein
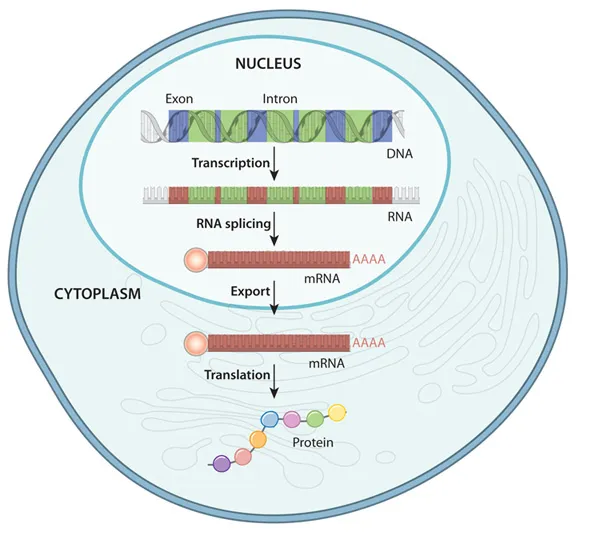
- RNA Processing (Eukaryotic Nucleus): Key modifications to pre-mRNA.
- 5' Capping: Addition of $m^7G$ (7-methylguanosine) cap; aids ribosome binding, protects mRNA from degradation.
- 3' Polyadenylation: Addition of poly-A tail (multiple adenine residues) to 3' end; enhances stability, facilitates nuclear export.
- Splicing: Removal of introns (non-coding regions) and joining of exons (coding regions) by spliceosome complex (snRNPs + proteins). Alternative splicing can produce multiple protein isoforms from a single gene.
- Translation (Cytoplasmic Ribosomes): Synthesis of protein from mRNA template.
- Genetic Code: Sequence of mRNA codons (triplets of nucleotides) determines amino acid sequence. AUG is the start codon (Methionine); UAA, UAG, UGA are stop codons. 📌 Mnemonic for Stop Codons: U Are Away (UAA), U Go Away (UGA), U Are Gone (UAG).
- Key Players:
- mRNA: Carries genetic code.
- tRNA: Adaptor molecule with anticodon (pairs with mRNA codon) and attached amino acid.
- Ribosomes: Composed of rRNA and proteins; site of protein synthesis (A, P, E sites).
- Steps: Initiation (assembly of ribosomal subunits, mRNA, initiator tRNA), Elongation (codon recognition, peptide bond formation, translocation), Termination (recognition of stop codon, release of polypeptide).
⭐ Wobble hypothesis: The base at the 5' end of the tRNA anticodon can form non-standard base pairs with the base at the 3rd position of the mRNA codon. This allows a single tRNA to recognize multiple codons for the same amino acid, reducing the number of tRNAs needed.
Gene Regulation - Who's the Boss?
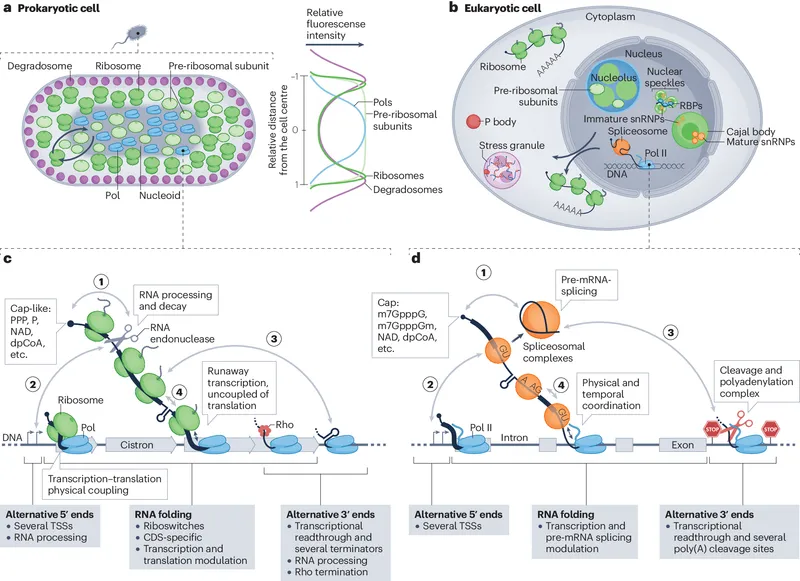
- Prokaryotes: Primarily at transcription.
- Operon Model: Coordinated gene clusters.
Feature Lac Operon (Catabolic) Trp Operon (Anabolic) Type Inducible (Default OFF) Repressible (Default ON) Inducer/Corep. Allolactose (inducer) Tryptophan (co-repressor) Repressor Active alone; inactivated by inducer Inactive alone; activated by co-repressor - Positive Control: e.g., CAP-cAMP in Lac operon.
- Operon Model: Coordinated gene clusters.
- Eukaryotes: Complex, multi-level control.
- Chromatin Remodeling: Histone Acetylation (↑ expression), Methylation (variable). 📌 HATs are HAppy (Active).
- Transcriptional: Transcription Factors (TFs), enhancers (↑), silencers (↓).
- Post-transcriptional: RNA processing (splicing, cap, tail), transport.
- Translational: miRNAs, siRNAs (RNAi) degrade/block mRNA.
- Post-translational: Folding, cleavage, modifications (e.g., phosphorylation).
⭐ Many eukaryotic genes are controlled by multiple enhancers and silencers, allowing for fine-tuned regulation in response to various signals.
High‑Yield Points - ⚡ Biggest Takeaways
- Lac operon exemplifies prokaryotic gene induction; Trp operon shows repression.
- Eukaryotic gene expression is controlled by transcription factors, enhancers/silencers, and chromatin structure.
- RNA Polymerase II is crucial for mRNA synthesis in eukaryotes.
- Post-transcriptional modifications include 5' capping, poly-A tail, and splicing of introns.
- The genetic code is degenerate, non-overlapping, and nearly universal.
- DNA methylation typically silences genes; histone acetylation often activates transcription.
- MicroRNAs (miRNAs) are small non-coding RNAs that regulate gene expression post-transcriptionally.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

