Basics of Gene Regulation - Control Central
- Gene Regulation: Microbes precisely control gene expression to adapt to environmental changes, conserve energy, and ensure cellular functions. Essential for survival and virulence.
- Levels of Control:
- Transcriptional: Most common; controls RNA synthesis.
- Translational: Controls protein synthesis from mRNA.
- Post-translational: Modifies protein activity after synthesis.
- Gene Types:
- Constitutive (Housekeeping): Always expressed; essential functions.
- Inducible/Repressible: Expression turned ON/OFF as needed.
- Key Elements:
- DNA Sequences: Promoters (initiate transcription), Operators (repressor binding site), Enhancers (activator binding site).
- Regulatory Proteins: Activators (↑ transcription), Repressors (↓ transcription).
⭐ Transcriptional control is the most energy-efficient and therefore the most common mode of gene regulation in prokaryotes.
The Operon Model - Switch On/Off
- Operon: Functional DNA unit: promoter, operator, structural genes.
- Lac Operon (Inducible - lactose catabolism)
- Genes: lacZ (β-galactosidase), lacY (permease), lacA (transacetylase).
- Negative Control: Repressor (LacI) binds operator (lactose absent). Inducer: Allolactose.
- Positive Control: ↓Glucose → ↑cAMP → CAP-cAMP complex binds promoter → ↑transcription.
- 📌 Mnemonic: LAG: Lac Absent, Gluc present=OFF; LAP: Lac Absent, Gluc Absent=OFF (CAP ready); LPG: Lac Present, Gluc present=Low expr; LPA: Lac Present, Gluc Absent=ON.
⭐ Allolactose, an isomer of lactose, is the actual inducer for the lac operon, not lactose itself.
- Trp Operon (Repressible - tryptophan synthesis)
- Genes: trpE, D, C, B, A.
- Negative Control: Repressor inactive. Corepressor: Tryptophan. High Trp → Repressor active → Operon OFF.
- Attenuation: Fine-tuning via leader sequence (trpL).
- Low Trp: Ribosome stalls → anti-termination loop → transcription ON.
- High Trp: Ribosome proceeds → termination loop → transcription OFF.

Broader Regulatory Networks - Smart Systems
- Catabolite Repression (Glucose Effect): Global regulation prioritizing glucose.
- Low Glucose: ↑ cAMP → cAMP binds Catabolite Activator Protein (CAP/CRP) → CAP-cAMP complex activates operons for alternative sugars (e.g., lac).
- High Glucose: ↓ cAMP → CAP inactive → repression of these operons.
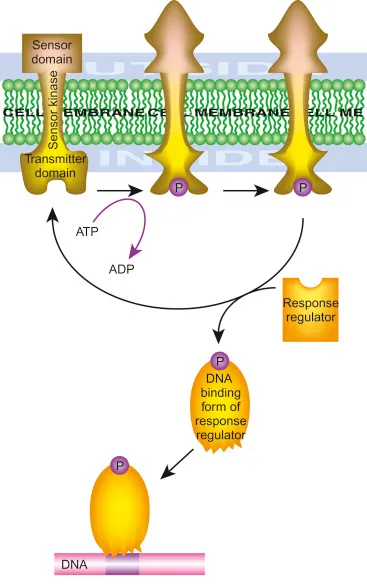
- Two-Component Regulatory Systems (TCS): Key for signal transduction.
- Sensor Kinase (SK): Transmembrane histidine kinase; detects external signal, autophosphorylates.
- Response Regulator (RR): Cytoplasmic; receives phosphate from SK, typically altering gene expression.
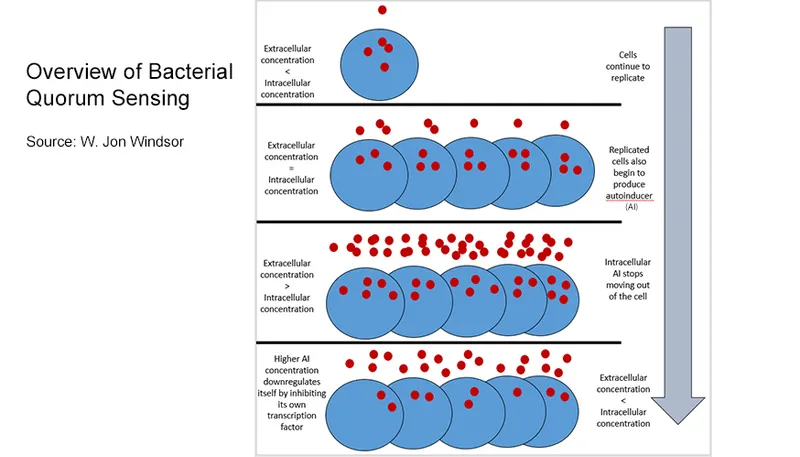
- Quorum Sensing (QS): Cell-density dependent gene regulation.
- Bacteria produce & release autoinducers (AIs).
- High cell density → ↑ AIs → threshold reached → coordinated gene expression.
- Autoinducers:
- Gram-negative: Acyl-Homoserine Lactones (AHLs).
- Gram-positive: Autoinducing Peptides (AIPs).
- Roles: Biofilm formation, virulence factor expression.
⭐ Many pathogenic bacteria utilize quorum sensing to coordinate the expression of virulence factors, only launching an attack when sufficient numbers are present.
High‑Yield Points - ⚡ Biggest Takeaways
- Operons (e.g., lac, trp) are fundamental for prokaryotic gene regulation.
- Lac operon: Inducible by lactose; CAP-cAMP mediates positive control (catabolite repression), repressor mediates negative control.
- Trp operon: Repressible by tryptophan; also features attenuation for fine-tuning.
- Sigma factors direct RNA polymerase to specific promoters for transcription initiation.
- Two-component regulatory systems allow bacteria to sense and adapt to environmental stimuli.
- Quorum sensing enables cell density-dependent gene expression and group behaviors.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

