Molecular Diagnostic Methods - PCR Powerhouse
- Polymerase Chain Reaction (PCR): In vitro exponential amplification of specific DNA/RNA.
- Core Principle: Enzymatic replication of a target nucleic acid sequence.
- Key Components: Template (DNA/cDNA), primers, Taq polymerase (heat-stable), dNTPs, Mg²⁺.
- PCR Cycle: Repeated 25-35 times. Results in $2^n$ fold amplification (n=cycles).
- Variants & Uses:
- RT-PCR (Reverse Transcriptase PCR): RNA detection (converts RNA to cDNA first). E.g., HIV, Influenza.
- qPCR (Real-Time PCR): Quantifies DNA/RNA during amplification using fluorescence.
- Applications: Rapid pathogen ID (TB, Chlamydia), genetic testing, oncology.
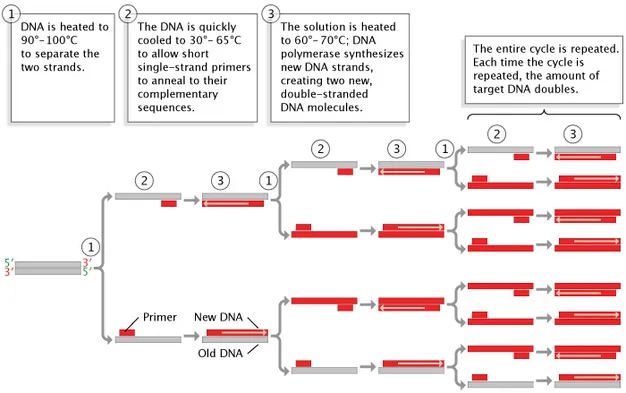
⭐ Taq polymerase, isolated from Thermus aquaticus, is thermostable, withstanding high denaturation temperatures, making PCR automation possible.
Molecular Diagnostic Methods - Target Finders
- Nucleic Acid Probes: Labelled (e.g., $P^{32}$, biotin, fluorophores) ssDNA/RNA fragments; bind specific complementary target sequences.
- Hybridization: Process of probe annealing to target nucleic acid.
- Solid-phase: Target immobilized on a membrane.
- In-situ: Target within intact cells or tissues.
- Blotting Techniques: For detecting specific DNA/RNA sequences.
- Southern Blot: DNA detection.
- **Northern Blot:** RNA detection (no restriction digestion needed).
- **Dot/Slot Blot:** Rapid detection without size separation.
> ⭐ 📌 SNoW DRoP Mnemonic: **S**outhern Blot = **D**NA; **N**orthern Blot = **R**NA; **W**estern Blot = **P**rotein.
- In Situ Hybridization (ISH): Localizes nucleic acids within cells/tissues.
- FISH (Fluorescent ISH): Uses fluorescent probes. Applications: Gene mapping, detecting aneuploidies (e.g., Trisomy 21), pathogen identification.
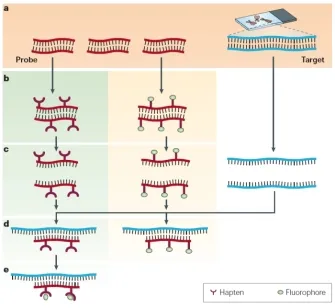 technique showing probes targeting specific DNA sequences in chromosomes)
technique showing probes targeting specific DNA sequences in chromosomes)
- FISH (Fluorescent ISH): Uses fluorescent probes. Applications: Gene mapping, detecting aneuploidies (e.g., Trisomy 21), pathogen identification.
- Microarrays (Gene Chips): Miniaturized arrays with thousands of probes; for high-throughput gene expression analysis, SNP genotyping.
Molecular Diagnostic Methods - Code Crackers
- Principle: "Crack the code" of microbial nucleic acids (DNA/RNA).
- Uses: ID, resistance, virulence, typing, epidemiology.

- 1. Hybridization (Known Sequences):
- Probes detect specific DNA/RNA.
- Techniques: Southern/Northern blots, FISH, DNA microarrays.
- 2. Amplification (Signal Boost):
- PCR: Exponential DNA/RNA increase. Key types: RT-PCR (RNA), qPCR (quantifies).
- Alternatives: LCR, TMA, NASBA.
- 3. Sequencing (Unknown Sequences):
- Sanger: Dideoxy chain termination. Gold standard for single genes.
- NGS: High-throughput, for genomes/metagenomes.
- 4. Genotyping/Typing (Strain Variation):
- RFLP, PFGE: Restriction enzyme patterns.
- MLST: Allelic variation in housekeeping genes. 📌 "Many Loci Sequenced Together".
⭐ PFGE is considered the gold standard for epidemiological typing of many bacterial pathogens.
Molecular Diagnostic Methods - Pathogen Hunt
Rapidly identifies pathogens by detecting their unique DNA/RNA, crucial for timely and targeted therapy.
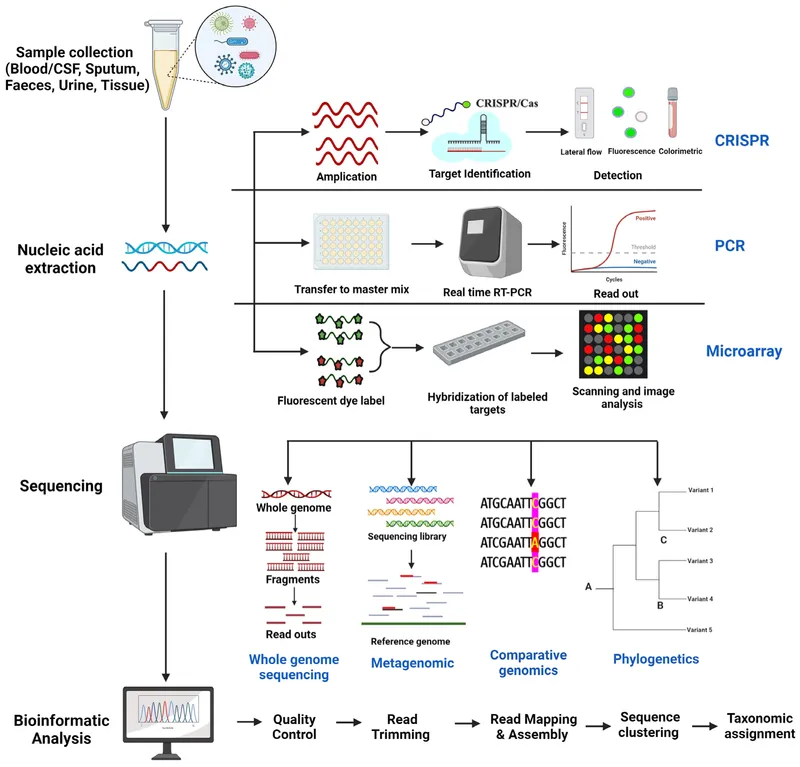
- Core Principle: Detect pathogen-specific nucleic acids.
- Key Methods:
- PCR (Polymerase Chain Reaction): Highly sensitive & specific.
- RT-PCR: For RNA viruses (e.g., HIV, Influenza, SARS-CoV-2).
- qPCR (Real-Time PCR): Quantifies pathogen load (viral load monitoring).
- Sequencing (NGS, Sanger): Definitive ID, tracks outbreaks, detects resistance mutations.
- LAMP (Loop-mediated Isothermal Amplification): Rapid, point-of-care testing.
- PCR (Polymerase Chain Reaction): Highly sensitive & specific.
- Advantages: Speed, detects non-culturable/fastidious organisms, high throughput.
- Applications: Diagnosis of sepsis, meningitis, TB, viral infections; antimicrobial resistance profiling.
⭐ PCR can detect as few as 1-10 copies of microbial DNA/RNA per sample, vital for early diagnosis even before seroconversion or in paucibacillary disease.
High‑Yield Points - ⚡ Biggest Takeaways
- PCR: Core for amplifying DNA/RNA; offers high sensitivity in detecting pathogens.
- Real-time PCR (qPCR): Allows quantification of microbial load, crucial for monitoring treatment response.
- Multiplex PCR: Detects multiple pathogens simultaneously, useful for syndromic panels.
- Nucleic Acid Hybridization Probes (e.g., FISH): Identify specific microbial sequences directly in samples.
- Gene Sequencing (Sanger/NGS): Essential for genotyping, resistance mutation detection, and epidemiological tracing.
- Molecular Typing (PFGE, MLST, Ribotyping): Crucial for strain differentiation during outbreak investigations.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

