Types of Mutations - Code's Quirky Typos
- Mutation: Permanent, heritable DNA sequence change.
- Point Mutations: Single base-pair (bp) change.
Type Change Consequence Example (Gene: AA change) Silent Codon → Same AA Usually no phenotypic effect Missense Codon → Different AA Variable; e.g., Sickle cell (HBB: Glu6Val) Sickle cell anemia Nonsense Codon → STOP (UAA, UAG, UGA) Truncated protein 📌 Stop Nonsense! β-thalassemia, Cystic Fibrosis Frameshift Indel (not multiple of 3 bp) Altered reading frame; new protein Tay-Sachs, Duchenne MD
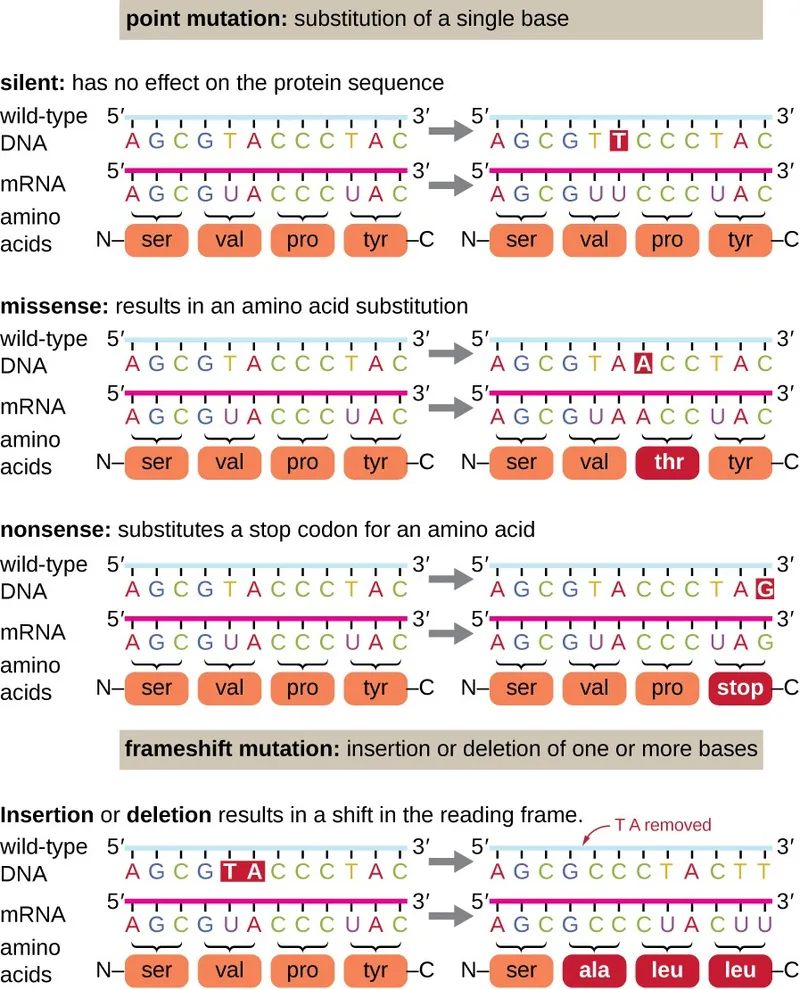
- Insertions/Deletions (Indels): Addition/removal of DNA bases; may cause frameshifts if not in multiples of 3 bp.
- Functional Effects:
- Loss-of-function (LOF): Reduced/null protein activity.
- Gain-of-function (GOF): New/enhanced protein activity.
- Dominant negative: Mutant protein antagonizes wild-type.
- Somatic vs. Germline:
- Somatic: In body cells; not inherited.
- Germline: In gametes; heritable.
⭐ Nonsense mutations cause premature stop codons, leading to truncated, often non-functional proteins.
Mutagens & Damage - DNA's Daily Dangers
DNA faces constant threats, leading to mutations if unrepaired.
Spontaneous Damage (Endogenous):
- Deamination: C → U; A → hypoxanthine.
- Depurination: Loss of purine base (A or G), creating an AP site.
- Oxidative Damage: Reactive oxygen species (ROS) modify bases (e.g., 8-oxoG).
Induced Damage (Exogenous Mutagens):
1. Chemical Mutagens:
- Base Analogs: e.g., 5-Bromouracil (5-BU) incorporates like T, mispairs with G.
- Alkylating Agents: e.g., Ethyl methanesulfonate (EMS) adds alkyl groups (ethyl to G), causing mispairing.
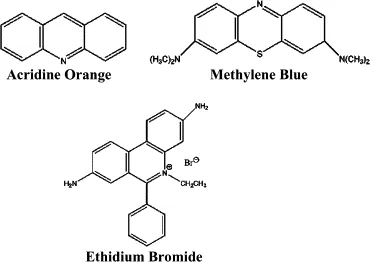
- Intercalating Agents: e.g., Ethidium bromide, proflavine. Insert between base pairs, causing frameshifts.
- Deaminating Agents: e.g., Nitrous acid ($HNO_2$) converts C→U, A→hypoxanthine.
2. Physical Mutagens:
- UV Radiation (Non-ionizing): Forms pyrimidine dimers (CPDs like T-T dimers) and 6-4 photoproducts.
- Ionizing Radiation (X-rays, γ-rays): Causes single-strand breaks (SSBs) and double-strand breaks (DSBs).
⭐ UV radiation predominantly causes cyclobutane pyrimidine dimers (CPDs) and 6-4 photoproducts (6-4PPs).
Single-Strand Repair - Precision Patchwork
-
Direct Repair: Directly reverses DNA damage.
- Photolyase (not in humans): Repairs pyrimidine dimers (UV) using light.
- MGMT ($O^6$-Methylguanine-DNA Methyltransferase): Removes alkyl groups from $O^6$-methylguanine. Suicide enzyme.
-
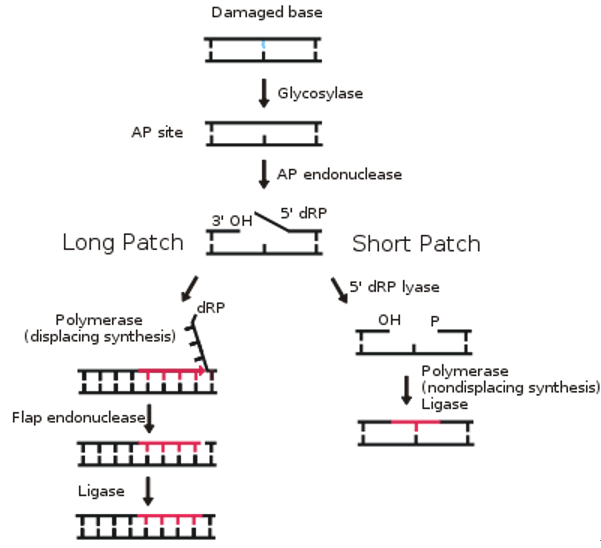
Base Excision Repair (BER): For small, non-helix distorting lesions (e.g., uracil, oxidized/alkylated bases).
- 📌 Mnemonic: GEL PLeASe (DNA Glycosylase, AP Endonuclease, AP Lyase, DNA Polymerase, DNA Ligase).
⭐ Base Excision Repair (BER) is initiated by a DNA glycosylase that recognizes and removes a specific damaged base, creating an AP site.
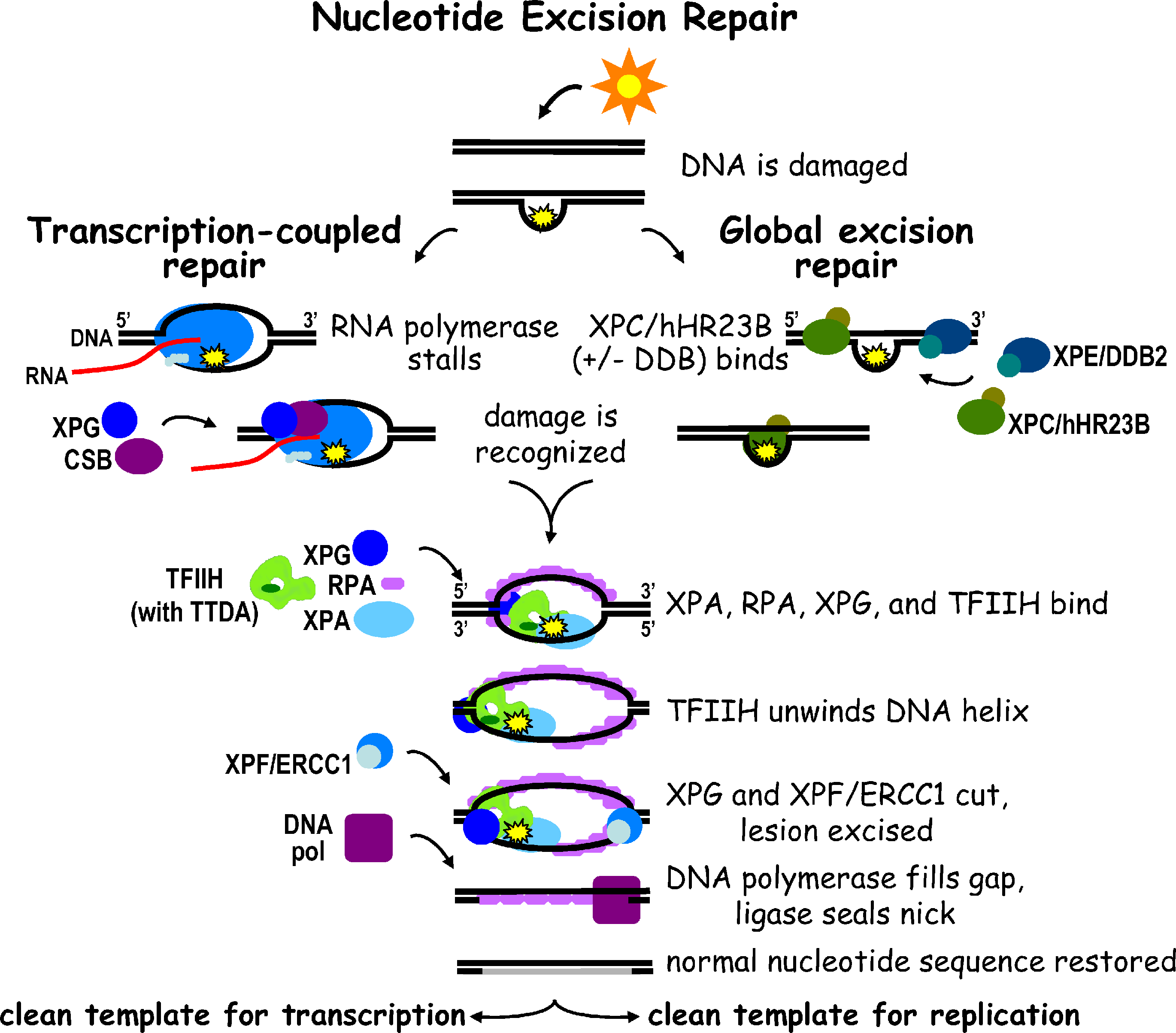
- Nucleotide Excision Repair (NER): For bulky, helix-distorting lesions (e.g., pyrimidine dimers, bulky adducts). Excises oligonucleotide (~24-32 nt in humans).
- Key Proteins:
- Humans: XP proteins (XPA-XPG, part of ~30 proteins). Deficiency → Xeroderma Pigmentosum.
- E. coli: Uvr proteins (UvrA,B,C excinuclease); excises ~12-13 nt.
- Key Proteins:
Complex Repair & Syndromes - When Fixes Falter
- Mismatch Repair (MMR): Corrects DNA replication errors.
- Prokaryotes: MutS, MutL, MutH.
- Eukaryotes: MSH, MLH, PMS.
- Double-Strand Break (DSB) Repair:
- NHEJ (Non-Homologous End Joining): Error-prone; predominant in G1.
- HR (Homologous Recombination): Error-free; S/G2 phase (uses sister chromatid). Proteins: BRCA1, BRCA2.
- Key Repair Defect Syndromes:
Syndrome Defective Pathway/Gene Key Features Xeroderma Pigmentosum NER UV sensitivity, ↑ skin cancer risk Lynch Syndrome (HNPCC) MMR ↑ colorectal, endometrial, ovarian cancer risk Ataxia Telangiectasia ATM (DSB sensing) Ataxia, telangiectasias, immunodeficiency, cancer BRCA1/BRCA2 Cancers HR ↑ breast, ovarian cancer risk Fanconi Anemia ICL repair, HR Pancytopenia, congenital abnormalities, cancer
⭐ Defects in Nucleotide Excision Repair (NER) are responsible for Xeroderma Pigmentosum, leading to extreme sensitivity to UV light and a high risk of skin cancer.
High‑Yield Points - ⚡ Biggest Takeaways
- Point mutations include silent, missense (altered amino acid), and nonsense (STOP codon).
- Frameshift mutations result from insertions/deletions not divisible by three, altering the reading frame.
- Mismatch Repair (MMR) corrects replication errors; defects lead to Lynch syndrome (HNPCC).
- Nucleotide Excision Repair (NER) fixes UV-induced pyrimidine dimers; defective in Xeroderma Pigmentosum.
- Base Excision Repair (BER) removes single damaged bases using DNA glycosylases.
- Double-strand breaks are repaired by NHEJ (error-prone) or Homologous Recombination (high-fidelity).
- The Ames test evaluates a chemical's mutagenic potential using bacteria_._
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more