Virology

On this page
🧬 Viral Architecture: The Molecular Invasion Machine
Viruses represent the ultimate biological minimalists-stripped-down genetic parasites that lack metabolic machinery and depend entirely on host cells for replication. Understanding viral structure reveals why certain antivirals work, how immune responses target infections, and what makes some viruses more pathogenic than others.
Every virus contains a nucleic acid core (DNA or RNA, never both) wrapped in a protein coat (capsid) composed of repeating capsomere subunits. This elegant design minimizes genetic baggage while maximizing infectivity. The capsid architecture determines antigenicity, stability, and transmission characteristics that define clinical syndromes.

Viral Genome Organization
-
DNA Viruses
- Double-stranded DNA (dsDNA): Herpesviruses, Adenoviruses, Poxviruses, Papillomaviruses, Polyomaviruses
- Genome size: 5-280 kb (largest in Poxviruses)
- Replication site: Nucleus (except Poxviruses-cytoplasm)
- Clinical significance: Latency potential in Herpesviruses
- Single-stranded DNA (ssDNA): Parvoviruses
- Genome size: 5 kb (smallest DNA viruses)
- Requires dividing cells (targets bone marrow, fetal tissues)
- Example: Parvovirus B19 causes aplastic crisis in sickle cell disease
- Double-stranded DNA (dsDNA): Herpesviruses, Adenoviruses, Poxviruses, Papillomaviruses, Polyomaviruses
-
RNA Viruses
- Positive-sense ssRNA (+RNA): Acts as mRNA, directly translated
- Picornaviruses, Flaviviruses, Togaviruses, Coronaviruses
- Infectious genome alone (no polymerase needed)
- >70% of human viral pathogens are RNA viruses
- Negative-sense ssRNA (−RNA): Requires viral polymerase
- Orthomyxoviruses, Paramyxoviruses, Rhabdoviruses, Filoviruses
- Must package RNA-dependent RNA polymerase in virion
- Higher mutation rate: 10⁻³ to 10⁻⁵ per nucleotide per replication
- Double-stranded RNA (dsRNA): Reoviruses, Rotaviruses
- Segmented genome: 10-12 segments
- Replication entirely in cytoplasm
- Retroviruses: +RNA with reverse transcriptase
- Convert RNA → DNA → integrate into host genome
- Error rate: 1 mutation per 10,000 nucleotides (HIV evolution driver)
- Positive-sense ssRNA (+RNA): Acts as mRNA, directly translated
📌 Remember: DNA viruses replicate in the Nucleus (except Pox-cytoplasm). RNA viruses replicate in the Cytoplasm (except Influenza and Retroviruses-nucleus for transcription/integration). Think "DNA Nucleus, RNA Cytoplasm" with 2 exceptions.
Capsid Symmetry and Clinical Implications
| Symmetry Type | Structure | Examples | Stability | Transmission | Clinical Pattern |
|---|---|---|---|---|---|
| Icosahedral | 20 triangular faces, 12 vertices | Adenovirus, Poliovirus, Papillomavirus | High environmental stability | Fecal-oral, respiratory, fomites | Epidemic outbreaks, seasonal peaks |
| Helical | Rod-shaped, nucleocapsid spiral | Influenza, Measles, Rabies | Lower stability (usually enveloped) | Respiratory droplets, animal bites | Requires close contact, zoonotic |
| Complex | Neither icosahedral nor helical | Poxviruses, Bacteriophages | Highest stability | Direct contact, aerosol | Bioterrorism concern (smallpox) |
⭐ Clinical Pearl: Icosahedral non-enveloped viruses survive gastric acid (pH 2-3), enabling fecal-oral transmission. Enveloped viruses are inactivated by bile salts, restricting them to respiratory/parenteral routes. This explains why Hepatitis A (non-enveloped) spreads through contaminated food while Hepatitis B/C (enveloped) require blood/body fluid exposure.
The Envelope: Lipid Armor and Achilles Heel
Enveloped viruses acquire a lipid bilayer from host cell membranes (plasma, nuclear, or ER) during budding. Embedded viral glycoproteins mediate attachment, fusion, and immune evasion.
-
Envelope Advantages
- Glycoprotein spikes enable receptor-specific cell targeting
- Membrane fusion allows direct cytoplasmic entry (bypassing endosomes)
- Antigenic variation through glycoprotein mutation (Influenza hemagglutinin)
- Complement evasion by incorporating host proteins
-
Envelope Vulnerabilities
- Desiccation sensitivity: Inactivated within minutes to hours outside host
- Detergent susceptibility: Soap disrupts lipid bilayer (hand hygiene critical)
- Alcohol sensitivity: 60-70% ethanol denatures envelope proteins
- Heat lability: Inactivated at 56°C for 30 minutes
💡 Master This: Enveloped viruses require close contact or body fluid transmission because environmental stability is poor. Non-enveloped viruses cause epidemic outbreaks via fomites and survive wastewater. When you see nosocomial clusters, think enveloped viruses (Influenza, RSV, Varicella). When you see community-wide epidemics, think non-enveloped (Norovirus, Adenovirus, Enteroviruses).
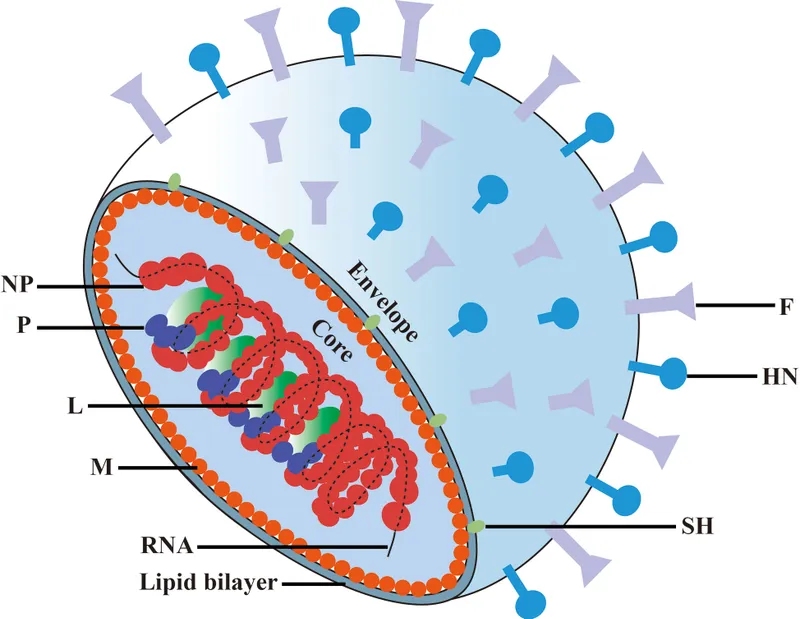
Viral Attachment Proteins: The Cellular Key
-
Hemagglutinin (HA) - Influenza
- Binds sialic acid receptors on respiratory epithelium
- 16 HA subtypes determine host tropism (avian vs human)
- Target of neutralizing antibodies (vaccine antigen)
-
gp120/gp41 - HIV
- gp120 binds CD4 + coreceptor (CCR5 or CXCR4)
- gp41 mediates membrane fusion
- CCR5-Δ32 mutation confers HIV resistance in 1% Europeans
-
Fiber protein - Adenovirus
- Binds CAR (coxsackie-adenovirus receptor)
- Penton base binds integrins → endocytosis
- Serotype-specific: >50 serotypes with different tropisms
-
VP1 protein - Poliovirus
- Binds CD155 (poliovirus receptor)
- Expressed on motor neurons → paralysis
- Receptor distribution determines neurotropism
📌 Remember: SHAFTED viruses have Surface Hemagglutinin And Fusion proteins To Enable Disease-Measles, Mumps, Parainfluenza, RSV, Influenza. These glycoproteins are diagnostic targets (hemagglutination assays) and vaccine antigens.
Connect viral structural features to their replication strategies through , where capsid uncoating and genome delivery mechanisms determine antiviral targeting opportunities.
🧬 Viral Architecture: The Molecular Invasion Machine
🔄 The Viral Replication Cycle: Hijacking Cellular Machinery
Viral replication follows a precisely orchestrated sequence that transforms host cells into viral production factories. Understanding each step reveals antiviral drug targets, explains viral tropism, and predicts clinical latency patterns. Every replication stage represents a potential therapeutic intervention point.
The replication cycle divides into 6 discrete phases: attachment → penetration → uncoating → biosynthesis → assembly → release. DNA and RNA viruses employ fundamentally different biosynthesis strategies, but all viruses face the same challenge: delivering genetic material into the host cell and hijacking cellular machinery for progeny production.
Attachment and Penetration: Gaining Cellular Entry
Viral attachment requires specific receptor-ligand interactions between viral surface proteins and host cell receptors. This molecular recognition determines tissue tropism, host range, and pathogenesis patterns.
-
Receptor-Mediated Endocytosis (most common)
- Virus binds receptor → clathrin-coated pit formation
- Endosome acidification (pH 5.0-6.0) triggers conformational changes
- Fusion proteins activated → viral membrane fuses with endosomal membrane
- Examples: Influenza (HA-mediated), Adenovirus, Poliovirus
- Amantadine blocks Influenza M2 ion channel, preventing uncoating
-
Direct Membrane Fusion (enveloped viruses)
- Fusion proteins insert into host plasma membrane at neutral pH
- Nucleocapsid delivered directly into cytoplasm
- Examples: HIV (gp41), Paramyxoviruses (F protein), HSV (gB/gD/gH/gL complex)
- Enfuvirtide blocks HIV gp41 fusion peptide
-
Pore Formation (non-enveloped viruses)
- Capsid proteins create membrane pores
- Genome translocates through pore into cytoplasm
- Examples: Poliovirus, Adenovirus (after endosomal escape)
⭐ Clinical Pearl: Viral receptor expression determines disease manifestations. Poliovirus binds CD155 on motor neurons (paralysis), Rabies binds nicotinic acetylcholine receptors on peripheral nerves (ascending CNS spread), EBV binds CD21 on B cells (infectious mononucleosis). Receptor distribution predicts clinical syndrome.

Uncoating: Genome Liberation Strategies
Uncoating releases viral genetic material from the protective capsid, making it accessible for replication. The location and mechanism vary by virus family.
-
Cytoplasmic Uncoating
- Picornaviruses: Receptor binding triggers conformational change → VP4 release → RNA injection
- Poxviruses: Two-stage uncoating (outer membrane removed at plasma membrane, core uncoated in cytoplasm)
- Timing: Within 30-60 minutes of penetration
-
Nuclear Pore Uncoating
- Herpesviruses: Capsid docks at nuclear pore complex → DNA injected through pore
- Adenoviruses: Partial uncoating in cytoplasm, complete uncoating at nuclear pore
- Requires intact microtubule network for capsid transport
-
Endosomal Uncoating
- Influenza: Low pH triggers HA conformational change → membrane fusion
- Requires M2 ion channel function (target of amantadine/rimantadine)
- Chloroquine raises endosomal pH, blocking uncoating (antiviral mechanism)
💡 Master This: Eclipse period-the interval between uncoating and new virion detection-ranges from 6-8 hours (Picornaviruses) to >24 hours (Herpesviruses). During eclipse phase, no infectious virus is detectable even though replication is active. This explains why antiviral therapy must start early; once assembly begins, intervention is less effective.
DNA Virus Biosynthesis: Nuclear Replication Strategies
DNA viruses (except Poxviruses) replicate in the nucleus using host DNA polymerases or encoding their own replication machinery.
-
Small DNA Viruses (Parvoviruses, Papillomaviruses, Polyomaviruses)
- Depend entirely on host S-phase machinery
- Parvovirus B19 requires rapidly dividing cells: bone marrow (aplastic crisis), fetal erythroblasts (hydrops fetalis)
- Papillomaviruses induce S-phase entry via E6/E7 oncoproteins
- Replication cycle: 12-24 hours
-
Large DNA Viruses (Herpesviruses, Adenoviruses)
- Encode own DNA polymerase, primase, helicase
- Independent of cell cycle (can infect non-dividing cells)
- Herpesvirus DNA pol: Target of acyclovir, ganciclovir, foscarnet
- Adenovirus: Rolling circle replication produces concatemers
- Replication cycle: 18-24 hours
-
Poxviruses (unique cytoplasmic DNA viruses)
- Encode complete replication machinery (>200 proteins)
- Cytoplasmic replication factories (viroplasm)
- Largest viral genomes: 130-280 kb
- Replication cycle: 12-14 hours
| DNA Virus Family | Replication Site | Host Machinery Dependence | DNA Polymerase | Replication Time | Antiviral Target |
|---|---|---|---|---|---|
| Parvovirus | Nucleus | Complete (S-phase required) | Host | 12-24 hours | None available |
| Papillomavirus | Nucleus | High (induces S-phase) | Host | 24-48 hours | Immune modulation |
| Adenovirus | Nucleus | Moderate | Viral | 18-24 hours | None specific |
| Herpesvirus | Nucleus | Low | Viral | 18-24 hours | Acyclovir, Ganciclovir |
| Poxvirus | Cytoplasm | None | Viral | 12-14 hours | Tecovirimat (ST-246) |
📌 Remember: Pox in the Box-Poxviruses replicate in the cytoplasmic "box" while all other DNA viruses need the nucleus. Poxviruses are the only DNA viruses replicating in cytoplasm because they encode complete transcription and replication machinery.
RNA Virus Biosynthesis: Diverse Replication Strategies
RNA viruses replicate in the cytoplasm (except Influenza and Retroviruses) using viral RNA-dependent RNA polymerase (RdRp). The lack of proofreading creates high mutation rates (10⁻³ to 10⁻⁵ per nucleotide per cycle).
-
Positive-Sense RNA Viruses (+RNA)
- Genome acts directly as mRNA → immediate translation
- First proteins synthesized: RdRp (replicase complex)
- RdRp creates (−) strand template → (+) strand progeny
- Examples: Poliovirus, Hepatitis C, Coronaviruses, Flaviviruses
- Replication cycle: 6-8 hours (fastest)
-
Negative-Sense RNA Viruses (−RNA)
- Genome cannot be translated → must package RdRp in virion
- RdRp creates (+) mRNA for translation
- (−) genome serves as template for (+) mRNA and (−) progeny
- Examples: Influenza, Measles, Mumps, Rabies, RSV
- Replication cycle: 8-12 hours
-
Double-Stranded RNA Viruses (dsRNA)
- Genome never fully uncoated (dsRNA triggers innate immunity)
- RdRp packaged in core transcribes mRNA through channels
- Examples: Rotavirus, Reovirus
- Segmented genomes enable reassortment (genetic shift)
- Replication cycle: 10-12 hours
-
Retroviruses (unique RNA → DNA strategy)
- Reverse transcriptase converts RNA → DNA
- Integrase inserts proviral DNA into host chromosome
- Host RNA pol II transcribes provirus → viral RNA
- Examples: HIV, HTLV
- Latency potential: Integrated provirus persists lifelong
- Replication cycle: 24-48 hours (integration adds time)
⭐ Clinical Pearl: Influenza is the only RNA virus requiring nuclear replication because it "cap-snatches"-steals 5' caps from host pre-mRNA to prime viral transcription. This nuclear dependence makes Influenza vulnerable to baloxavir marboxil, which inhibits cap-snatching endonuclease. All other (−)RNA viruses replicate entirely in cytoplasm.

💡 Master This: RNA virus mutation rates drive antigenic variation and antiviral resistance. HIV reverse transcriptase lacks proofreading, generating 1 mutation per genome per replication cycle. In untreated patients with viremia of 10¹⁰ virions/day, every possible single-point mutation occurs daily. This explains why HIV monotherapy fails within weeks-resistant mutants pre-exist. Combination antiretroviral therapy (cART) prevents resistance by requiring simultaneous mutations in multiple genes (probability <10⁻¹⁴).
Assembly and Release: Progeny Virus Liberation
Viral components converge at assembly sites where nucleocapsids form and acquire infectivity. Release mechanisms determine whether host cell survives infection.
-
Lytic Release (non-enveloped viruses)
- Virion accumulation → cell lysis → simultaneous release of 10,000-100,000 virions
- Examples: Adenovirus, Poliovirus, Papillomavirus
- Burst size: Number of progeny per infected cell
- Clinical: Acute cytopathic effect, tissue necrosis
-
Budding Release (enveloped viruses)
- Viral proteins insert into host membranes → nucleocapsid buds through
- Acquires lipid envelope during egress
- Cell may survive and continue producing virions (persistent infection)
- Examples: Influenza, HIV, Herpesviruses, Paramyxoviruses
- Neuraminidase (Influenza) cleaves sialic acid to release budding virions-target of oseltamivir/zanamivir
-
Exocytosis (some enveloped viruses)
- Virions bud into ER/Golgi → transported in vesicles → exocytosed
- Examples: Hepatitis B, Bunyaviruses
- Cell survival enables chronic infection
📌 Remember: LEMON viruses cause Lytic, Enveloped, Membrane-damaging, Oncogenic, Necrotic infections-but actually, this mnemonic is backwards! Non-enveloped viruses cause lytic release (Adenovirus, Poliovirus, Parvovirus). Enveloped viruses bud without immediate lysis (HIV, Influenza, Herpesviruses). Think "Envelope = Escape without Explosion."
Viral replication strategies directly influence pathogenesis patterns explored in , where replication kinetics determine acute versus chronic disease manifestations.
🔄 The Viral Replication Cycle: Hijacking Cellular Machinery
🧪 Viral Pathogenesis: From Infection to Disease
Understanding how viruses cause disease requires integrating viral replication with host immune responses, tissue tropism, and damage mechanisms. Pathogenesis determines clinical presentation, diagnostic approach, and therapeutic intervention timing. The severity of viral disease reflects the balance between viral virulence factors and host defense mechanisms.
Viral pathogenesis unfolds through predictable stages: entry → local replication → dissemination → target organ infection → immune response → resolution or chronicity. Each stage presents distinct clinical features and intervention opportunities.
Entry Portals and Primary Replication
Viruses enter through 4 major portals: respiratory tract (most common, >60% of infections), gastrointestinal tract, skin/mucosa, and parenteral routes. Portal characteristics determine which viruses successfully establish infection.
-
Respiratory Tract Entry
- Aerosol droplets: 5-10 μm particles reach lower airways (Influenza, Measles, SARS-CoV-2)
- Larger droplets: >10 μm deposit in upper airways (Rhinovirus, Adenovirus)
- Mucociliary clearance: 1 cm/hour escalator removes trapped virions
- Alveolar macrophages: First-line defense, phagocytose >90% of inhaled particles
- IgA in respiratory secretions: 2-5 mg/mL neutralizes viruses at mucosal surface
-
Gastrointestinal Tract Entry
- Must survive gastric acid (pH 2-3), bile salts, proteases
- Only non-enveloped viruses: Enteroviruses, Rotavirus, Norovirus, Hepatitis A
- M cells in Peyer's patches: Transcytose viruses across intestinal epithelium
- Primary replication in intestinal lymphoid tissue
- Fecal shedding: 10⁸-10¹¹ virions/gram (high transmissibility)
-
Skin and Mucosal Entry
- Requires epithelial breach: Arthropod bites (Arboviruses), needlestick (HIV, HBV, HCV)
- Sexual transmission: Microabrasions in genital mucosa (HSV, HIV, HPV)
- Langerhans cells: Dendritic cells capture virus → transport to lymph nodes
- Vertical transmission: Transplacental (CMV, Rubella, Parvovirus B19)
⭐ Clinical Pearl: Incubation period correlates with distance from entry site to target organ. Short incubation (<7 days): Local replication only (Influenza, Rhinovirus, Rotavirus). Medium incubation (7-21 days): Viremia required (Measles, Mumps, Varicella). Long incubation (>21 days): Neural spread or slow replication (Rabies 30-90 days, HIV weeks to months). Incubation period guides post-exposure prophylaxis timing.

Viral Dissemination: Local Versus Systemic Spread
After primary replication at entry site, viruses either remain localized or disseminate systemically via blood or nerves.
-
Localized Infections (no viremia)
- Replication confined to entry portal
- Examples: Rhinovirus (upper respiratory), Rotavirus (intestinal epithelium), Papillomavirus (skin/mucosa)
- Short incubation: 1-3 days
- Symptoms correlate directly with local tissue damage
- Immunity: Mucosal IgA predominates
-
Systemic Infections with Viremia
- Primary viremia: Initial spread from entry site to reticuloendothelial system (RES)
- Magnitude: 10²-10⁴ virions/mL
- Duration: 2-3 days
- Often asymptomatic
- Secondary viremia: Amplification in RES → massive bloodstream release
- Magnitude: 10⁴-10⁸ virions/mL
- Duration: 3-7 days
- Correlates with symptom onset (fever, rash, systemic symptoms)
- Examples: Measles, Varicella, Enteroviruses, Hepatitis viruses
- Primary viremia: Initial spread from entry site to reticuloendothelial system (RES)
-
Neural Spread (neurotropic viruses)
- Retrograde axonal transport: 2-3 mm/hour along peripheral nerves
- Examples: Rabies (from bite site to CNS), HSV (sensory ganglia), VZV (dorsal root ganglia)
- Establishes latency in neurons (immune-privileged site)
- Reactivation: Anterograde transport to skin → vesicular lesions
💡 Master This: Viremia pattern predicts rash timing and distribution. Centrifugal rash (face → trunk → extremities) indicates hematogenous dissemination with uniform seeding (Measles, Rubella). Centripetal rash (extremities → trunk) suggests sequential local replication (Varicella-new crops appear as virus spreads). Rash appears 12-14 days after exposure in viremic exanthems, coinciding with peak secondary viremia.
Tissue Tropism: Viral Target Preferences
Tissue tropism-the preferential infection of specific cell types-determines clinical syndrome and disease severity. Tropism reflects receptor expression, intracellular permissiveness, and immune surveillance.
-
Receptor Expression
- HIV: CD4 + CCR5/CXCR4 on T-helper cells, macrophages, microglia
- Poliovirus: CD155 on motor neurons (anterior horn cells)
- EBV: CD21 (CR2) on B cells and nasopharyngeal epithelium
- Rabies: Nicotinic acetylcholine receptor on neurons
- Measles: CD46 and SLAM on lymphocytes, respiratory epithelium
-
Intracellular Permissiveness
- Cell must provide replication machinery (S-phase for Parvoviruses)
- Interferon sensitivity: Hepatocytes produce robust IFN response (limits HCV replication)
- Protease availability: HIV requires CD4+ T cell activation for productive infection
-
Immune Privilege
- CNS: Blood-brain barrier limits antibody access, neurons don't express MHC-I
- Eye: Anterior chamber-associated immune deviation (ACAID)
- Testes: Sertoli cell barrier protects from immune attack
- Viruses exploit these sites for persistence: HSV (neurons), CMV (retina), Mumps (testes)
| Virus | Primary Target Cells | Key Receptor | Clinical Manifestation | Tropism Determinant |
|---|---|---|---|---|
| HIV | CD4+ T cells, Macrophages | CD4 + CCR5/CXCR4 | Immunodeficiency | Coreceptor expression |
| Influenza | Respiratory epithelium | Sialic acid (α2,6 human, α2,3 avian) | Pneumonia | Hemagglutinin specificity |
| Hepatitis B | Hepatocytes | NTCP (sodium taurocholate transporter) | Hepatitis, Cirrhosis | Liver-specific transporter |
| Poliovirus | Motor neurons | CD155 (PVR) | Flaccid paralysis | Neuron-specific receptor |
| Measles | Lymphocytes, Respiratory epithelium | CD46, SLAM (CD150) | Rash, Pneumonia, Encephalitis | Immune cell targeting |
| Rabies | Neurons | Nicotinic ACh receptor | Encephalitis | Neurotropism |
📌 Remember: HEAVEN viruses cause Hepatotropic, Encephalotropic, Arthritogenic, Viremic, Enterotropic, Neurotropic infections based on receptor distribution. Hepatitis viruses (A, B, C, D, E) target liver via specific transporters. Enteroviruses (Polio, Coxsackie, Echo) target intestinal M cells. Neurotropic viruses (Rabies, HSV, VZV, JC virus) infect neurons. Tropism predicts clinical syndrome.
Mechanisms of Viral Tissue Damage
Viruses damage tissues through direct cytopathic effects and immune-mediated pathology. The relative contribution determines disease severity and therapeutic approach.
-
Direct Cytopathic Effects
- Cell lysis: Poliovirus, Adenovirus, Influenza (release 10,000-100,000 virions per cell)
- Syncytia formation: RSV, Measles, HSV, HIV (fusion of infected cells → multinucleated giant cells)
- Inclusion bodies: Intranuclear (Herpesviruses-Cowdry type A) or cytoplasmic (Rabies-Negri bodies)
- Apoptosis induction: HIV, Adenovirus (trigger programmed cell death)
- Metabolic disruption: Poliovirus shuts off host protein synthesis within 30 minutes
-
Immune-Mediated Pathology
- CTL-mediated killing: Hepatitis B liver damage primarily from CD8+ T cell attack on infected hepatocytes
- Antibody-dependent enhancement: Dengue secondary infection with heterologous serotype
- Immune complex deposition: Hepatitis B causing glomerulonephritis, polyarteritis nodosa
- Cytokine storm: Influenza H5N1, SARS-CoV-2 severe disease (IL-6 >100 pg/mL)
- Autoimmunity: Molecular mimicry (Coxsackie B → myocarditis via cardiac myosin cross-reactivity)
⭐ Clinical Pearl: Hepatitis B severity paradox-immunocompetent patients develop acute hepatitis (immune-mediated damage), while immunocompromised patients remain asymptomatic carriers (minimal CTL response). Chronic HBV carriers have 10⁵-10⁸ virions/mL but normal transaminases until immune reconstitution triggers hepatitis. This explains why immunosuppression withdrawal causes hepatitis flares.
Connect viral pathogenesis mechanisms to specific viral families through , where detailed immune evasion strategies and persistence mechanisms are explored.
🧪 Viral Pathogenesis: From Infection to Disease
🦠 Oncogenic Viruses: Molecular Carcinogenesis
Approximately 15-20% of human cancers worldwide are attributable to viral infections. Oncogenic viruses transform cells through sustained expression of viral oncoproteins that dysregulate cell cycle checkpoints, inhibit tumor suppressors, and promote genomic instability. Understanding viral oncogenesis mechanisms reveals cancer prevention strategies (vaccination) and therapeutic targets.
Oncogenic viruses fall into DNA virus and RNA virus (retrovirus) categories, each employing distinct transformation mechanisms. DNA viruses typically encode oncoproteins that inactivate p53 and Rb tumor suppressors, while retroviruses integrate into host genomes and activate proto-oncogenes or encode viral oncogenes.
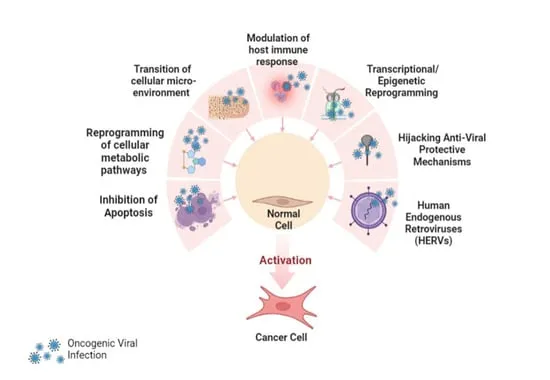
DNA Oncogenic Viruses: Cell Cycle Hijackers
-
Human Papillomaviruses (HPV)
- High-risk types: HPV-16, 18 (cause 70% of cervical cancers worldwide)
- Oncoproteins: E6 and E7
- E6: Binds p53 → ubiquitin-mediated degradation → loss of apoptosis
- E7: Binds Rb → releases E2F transcription factors → uncontrolled S-phase entry
- Combined effect: Immortalization and genomic instability
- Cancers: Cervical (99% HPV+), oropharyngeal (70% HPV+), anal, vulvar, penile
- Integration into host genome: Disrupts E2 repressor → constitutive E6/E7 expression
- Latency period: 10-30 years from infection to cancer
- Prevention: HPV vaccine (Gardasil 9) covers 9 types (6, 11, 16, 18, 31, 33, 45, 52, 58)
-
Epstein-Barr Virus (EBV)
- Infects >90% of adults worldwide (usually asymptomatic or infectious mononucleosis)
- Oncoproteins: LMP1, LMP2, EBNA1, EBNA2
- LMP1: Mimics constitutively active CD40 → NF-κB activation → proliferation
- EBNA2: Transactivates c-myc and other growth genes
- Cancers:
- Burkitt lymphoma (endemic African form: 100% EBV+, c-myc translocation t(8;14))
- Nasopharyngeal carcinoma (100% EBV+ in endemic regions-Southern China)
- Hodgkin lymphoma (40% EBV+, Reed-Sternberg cells)
- Post-transplant lymphoproliferative disorder (PTLD) in immunosuppressed patients
- Latency types: Type III (all oncoproteins expressed-PTLD) vs Type II (LMP1/2 only-Hodgkin) vs Type I (EBNA1 only-Burkitt)
-
Hepatitis B Virus (HBV)
- Chronic HBV infection increases hepatocellular carcinoma (HCC) risk 100-fold
- Mechanisms:
- Chronic inflammation → oxidative DNA damage → mutations
- HBx protein: Transactivates growth-promoting genes, inactivates p53
- Viral integration: Disrupts tumor suppressor genes, activates telomerase
- HCC develops in 3-5% of chronic carriers over 20-30 years
- Cirrhosis accelerates: Annual HCC incidence 3-8% in cirrhotic patients
- Prevention: HBV vaccination reduces HCC incidence by >70% (Taiwan experience)
-
Kaposi Sarcoma-Associated Herpesvirus (KSHV/HHV-8)
- Oncoproteins: LANA, v-cyclin, v-FLIP, v-IL-6
- LANA: Binds and inactivates p53 and Rb
- v-cyclin: Drives cell cycle progression, resistant to CDK inhibitors
- Cancers: Kaposi sarcoma (AIDS-defining, >90% KSHV+), primary effusion lymphoma, Castleman disease
- Endemic in Mediterranean and sub-Saharan Africa (5-40% seroprevalence)
- AIDS-associated KS: 10,000-fold increased risk in HIV+ KSHV+ individuals
- Oncoproteins: LANA, v-cyclin, v-FLIP, v-IL-6
| Oncogenic DNA Virus | Oncoproteins | Tumor Suppressor Targets | Associated Cancers | Latency Period | Prevention |
|---|---|---|---|---|---|
| HPV-16/18 | E6, E7 | p53 (E6), Rb (E7) | Cervical, Oropharyngeal | 10-30 years | HPV vaccine |
| EBV | LMP1, EBNA2 | Apoptosis inhibition | Burkitt, Nasopharyngeal, Hodgkin | 5-40 years | None available |
| HBV | HBx | p53 inactivation | Hepatocellular carcinoma | 20-30 years | HBV vaccine |
| KSHV (HHV-8) | LANA, v-cyclin | p53, Rb | Kaposi sarcoma | Variable (AIDS-accelerated) | ART for HIV+ |
📌 Remember: HEPK DNA viruses are Highly Efficient at Producing Kancer-HPV, EBV, Polyomavirus (rare), KSHV. All encode proteins targeting p53/Rb pathways. Think "DNA viruses Delete Natural Apoptosis."
RNA Oncogenic Viruses: Retroviral Integration
-
Human T-Lymphotropic Virus Type 1 (HTLV-1)
- Endemic in Japan, Caribbean, Central Africa (10-20 million infected worldwide)
- Mechanism: Tax oncoprotein
- Transactivates NF-κB, CREB, AP-1 → T-cell proliferation
- Inactivates p53, inhibits DNA repair
- Induces IL-2 and IL-2 receptor → autocrine growth loop
- Cancer: Adult T-cell leukemia/lymphoma (ATL)
- Develops in 2-5% of HTLV-1 carriers
- Latency: 20-40 years (longest of any oncogenic virus)
- Aggressive clinical course: Median survival 6-10 months
- Also causes: Tropical spastic paraparesis (HAM/TSP)-1-2% of carriers
-
Human Immunodeficiency Virus (HIV)
- Indirect oncogenesis via immunosuppression
- Allows oncogenic virus reactivation: KSHV (Kaposi sarcoma), EBV (lymphomas), HPV (cervical/anal cancer)
- AIDS-defining cancers: Kaposi sarcoma, non-Hodgkin lymphoma, cervical cancer
- Non-AIDS-defining cancers: Increased risk of lung, liver, anal, Hodgkin lymphoma
- CD4 count <200 cells/μL: Highest cancer risk (10-20 fold increased)
- ART reduces but doesn't eliminate cancer risk (chronic inflammation persists)
⭐ Clinical Pearl: Cervical cancer screening in HIV+ women requires more frequent intervals-annual Pap smears versus every 3 years in HIV-negative women. HPV persistence is 3-5 times more common in HIV+ women, and progression to high-grade dysplasia occurs 2-3 times faster. This reflects impaired cell-mediated immunity against HPV-infected cells.
💡 Master This: Oncogenic virus vaccination represents the most successful cancer prevention strategy. HBV vaccination in Taiwan reduced childhood HCC incidence by 70% within 20 years. HPV vaccination is projected to prevent >90% of cervical cancers if administered before sexual debut. No other
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app














