PCR: Core Principle - Tiny DNA Photocopying
- Polymerase Chain Reaction (PCR): An in-vitro technique for exponential amplification of a target DNA segment.
- Invented by Kary Mullis (Nobel Prize, 1993).
- Acts as a "molecular photocopier," creating millions to billions of DNA copies from a small initial amount.
- This selective amplification allows for subsequent analysis, even with minimal starting material.
- Fundamental to molecular diagnostics, genetics, and forensics.
⭐ PCR can amplify a specific DNA sequence over a billion-fold in a few hours, starting from even a single DNA molecule or cell.
PCR: Essential Components - The Reaction Cocktail
📌 Mnemonic: "Daring Primates Take Nice Green Bananas & Magnesium" (DNA template, Primers, Taq Polymerase, dNTPs, Buffer, $MgCl_2$)
Key components and their roles:
| Component | Role |
|---|---|
| DNA Template | Source DNA; contains target sequence for amplification |
| Primers (Fwd/Rev) | Short oligos; define target region; initiate synthesis |
| Taq Polymerase | Thermostable DNA polymerase; synthesizes new DNA |
| dNTPs (A,T,C,G) | Deoxynucleotide triphosphates; DNA building blocks |
| Buffer | Maintains optimal pH for Taq polymerase activity |
| $MgCl_2$ | Cofactor for Taq; crucial for primer annealing, enzyme activity |
⭐ $MgCl_2$ concentration is critical: too low ↓ yield; too high ↑ non-specific products. Affects primer annealing & enzyme fidelity.
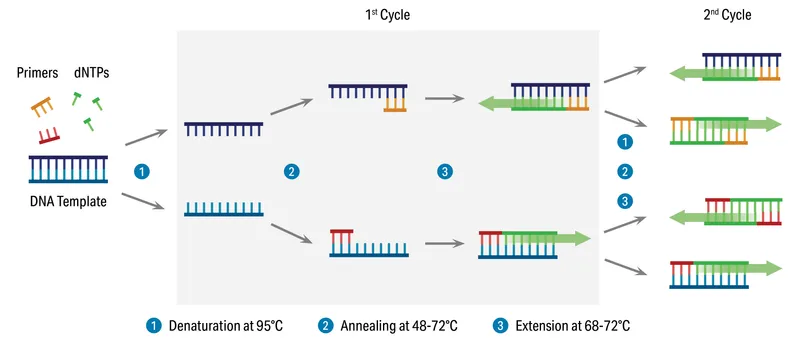
PCR: The Amplification Cycle - DNA Heat Dance
- The "heat dance": A precise sequence of temperature changes. Each cycle doubles target DNA.
- 📌 Mnemonic: DAE - Denature, Anneal, Extend.
- Denaturation: 94-98°C. Heat separates dsDNA into two single strands (ssDNA).
- Annealing: 50-65°C. Primers (short DNA sequences) bind to complementary sites on ssDNA.
- Extension: 72°C. Heat-stable Taq polymerase extends primers, synthesizing new DNA strands.
- Typically 25-35 cycles. Amplification formula: $2^n$ (n = number of cycles).
⭐ Magnesium ions (Mg²⁺) act as a cofactor for Taq polymerase and their concentration is critical for PCR efficiency and specificity.
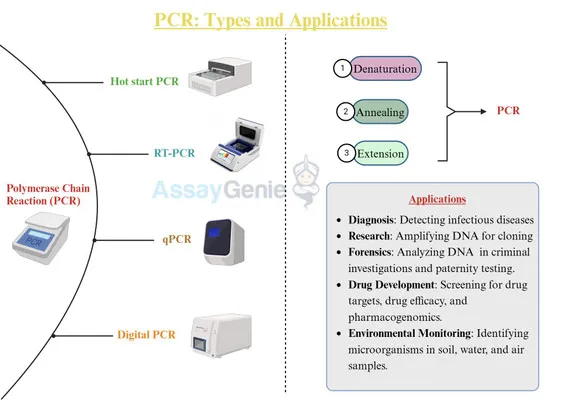
PCR: Key Variants - PCR's Special Moves
- RT-PCR (Reverse Transcriptase PCR): RNA → cDNA. Detects RNA viruses (HIV, SARS-CoV-2), gene expression.
- qPCR (Quantitative/Real-Time PCR): Real-time DNA/cDNA quantification. For pathogen load, gene expression.
- Nested PCR: Two PCR rounds; outer then inner primers. ↑Sensitivity/specificity. Detects low-abundance targets.
- Multiplex PCR: Multiple primer sets, one reaction. Detects several targets simultaneously (pathogen panels).
📌 PCR Variants: Really Quick, Neatly Multiplexed! (RT, qPCR, Nested, Multiplex)
⭐ qPCR (Real-Time PCR) allows for quantification of nucleic acids using fluorescent probes like TaqMan or dyes like SYBR Green.
PCR: Clinical Utility & Caveats - Microbe Detective Tool
- Clinical Utility:
- Rapid diagnosis: Viral (HIV), bacterial (TB), fungal infections.
- Detects non-culturable or slow-growing microbes.
- Quantifies microbial load (e.g., viral load).
- Identifies antimicrobial resistance genes.
- Epidemiological tracking & genotyping.
- Caveats:
- High sensitivity: Risk of false positives via contamination.
- Detects DNA/RNA, not necessarily live pathogens.
- Sample inhibitors can cause false negatives.
- Requires known target genetic sequence.
⭐ PCR's high sensitivity allows detecting pathogens in low numbers, often before symptoms fully develop.
High‑Yield Points - ⚡ Biggest Takeaways
- PCR achieves exponential amplification of target DNA sequences in vitro.
- Key requirements: Taq polymerase (thermostable), primers (short DNA sequences), dNTPs, template DNA.
- Cycles through Denaturation (
95°C), Annealing (primer-specific temp), Extension (72°C).- RT-PCR detects RNA (e.g., viruses) by first creating cDNA using reverse transcriptase.
- qPCR (Real-Time PCR) quantifies nucleic acids by monitoring fluorescence during amplification.
- Crucial for diagnosing infections, genetic disorders, and in forensic science.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

