Metagenomics Fundamentals - Unmasking Microbial Worlds
- Metagenomics: Culture-independent study of collective genomes from a microbial community, revealing its composition and functional potential.
| Feature | Metagenomics | Traditional Culturing |
|---|---|---|
| Principle | Direct genetic analysis of entire community | Isolation & lab growth of microbes |
| Scope | All microbes (culturable & unculturable) | Only culturable microbes (often <1%) |
| Data Yield | Genomic sequences, functional gene annotation | Phenotypic traits, limited genetic data |
| Discovery | High for novel genes, pathways, & species | Limited to known, culturable organisms |
Metagenomics Workflow - From Mud to Data
- Analyzes collective microbial genomes directly from an environmental sample.
- Sample Collection: Representative sample collection; immediate nucleic acid preservation.
- DNA/RNA Extraction: Efficient lysis of diverse microbes; DNA/RNA purification; inhibitor removal.
- Library Preparation: DNA fragmentation, adapter ligation, size selection.
- Sequencing Approaches:
- Shotgun metagenomics: Whole genome sequencing; reveals functions, novel genes.
- Amplicon sequencing: Marker gene (e.g., 16S rRNA, ITS) sequencing for taxonomy.
- Bioinformatic Processing:
- Read Quality Control (QC), filtering & trimming.
- Assembly (shotgun): Contig/Metagenome-Assembled Genome (MAG) reconstruction.
- Binning: Group sequences into Operational Taxonomic Units (OTUs) or MAGs.
- Functional (e.g., KEGG) & taxonomic (e.g., SILVA) annotation.
- Data Analysis: Ecological diversity ($α$, $β$), comparative genomics, pathway reconstruction.
⭐ Metagenome-Assembled Genomes (MAGs) enable genomic study of uncultured microbes, expanding known biodiversity & function.
Sequencing & Analysis - Decoding the Data Deluge
-
Predominantly uses Next-Generation Sequencing (NGS) (e.g., Illumina).
-
Two Main Approaches:
Feature Shotgun Metagenomics Amplicon (e.g., 16S rRNA) Sequencing Target Entire genomic DNA Specific marker genes (e.g., 16S, ITS) Output "Who is there?" & "What can they do?" "Who is there?" (Taxonomy) Discovery Potential High (genes, organisms) Low to moderate Host DNA Impact Significant interference possible Less affected (primer-dependent) Cost & Complexity Higher Lower -
Bioinformatic Pipeline Stages:
- Data Pre-processing: Quality control (QC), adapter trimming, host DNA removal.
- Assembly (Shotgun): Reconstructing genomes from reads (contigs, scaffolds).
- Taxonomic Profiling/Binning: Assigning sequences to taxa (e.g., OTUs, ASVs).
- Functional Annotation: Predicting gene functions and metabolic pathways.
- Statistical Analysis: Diversity indices (alpha, beta), comparative analysis.
⭐ The 16S rRNA gene is the most common molecular marker for bacterial and archaeal taxonomic profiling in amplicon sequencing due to its conserved and variable regions, crucial for species identification and phylogenetic studies.
Clinical Metagenomics - Microbes in Medicine
- Culture-independent microbial DNA/RNA analysis from patient samples (e.g., blood, CSF, stool).
- Key Applications:
- Diagnosing elusive or polymicrobial infections (e.g., sepsis, encephalitis).
- Rapidly detecting antimicrobial resistance (AMR) genes.
- Analyzing microbiome dysbiosis in diseases (IBD, obesity, cancer).
- Tracking outbreaks & pathogen surveillance.
- Guiding personalized antimicrobial therapy.
- Methods: Primarily Next-Generation Sequencing (NGS) - common approaches include 16S rRNA gene sequencing and shotgun metagenomics.
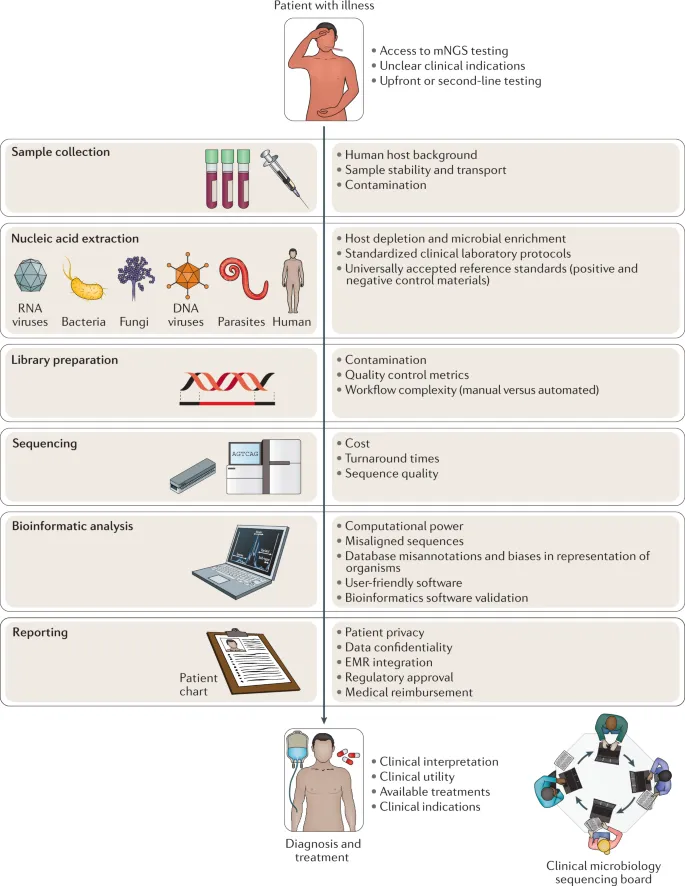
⭐ Shotgun metagenomics enables simultaneous detection of bacteria, viruses, fungi, and parasites from a single clinical sample, offering a comprehensive diagnostic view.
- Challenges: Cost, bioinformatics complexity, host DNA contamination, result interpretation for clinical actionability.
High‑Yield Points - ⚡ Biggest Takeaways
- Metagenomics: Culture-independent analysis of total genetic material from complex samples.
- Unveils microbial diversity, especially unculturable organisms (the "microbial dark matter").
- Identifies novel genes, enzymes, and antibiotic resistance determinants.
- Key for microbiome studies (e.g., gut dysbiosis) and pathogen discovery.
- Relies heavily on Next-Generation Sequencing (NGS) and bioinformatic analysis.
- 16S rRNA gene sequencing is widely used for bacterial and archaeal profiling.
- Helps understand community function and host-microbe interactions in health and disease.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

