IS Elements & Basics - Jumping Gene Intro
- Transposable Elements (TEs) or "jumping genes": DNA segments that can move (transpose) from one genomic location to another.
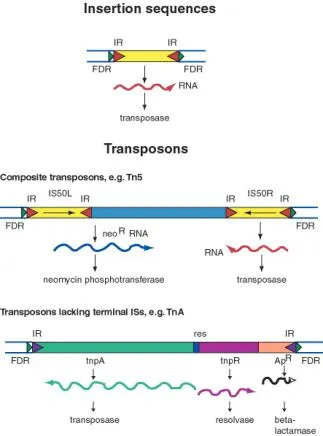
- Insertion Sequences (IS elements): Simplest TEs, acting as mobile genetic elements in bacteria.
- Size: Typically 0.7-2.5 kb.
- Structure: Contain a single open reading frame (ORF) encoding transposase, the enzyme essential for their movement.
- Flanked by short Inverted Repeats (IRs) at their ends; these sequences are recognized by transposase.
- Transposase: Mediates the excision and insertion of the IS element.
- Impact: Insertion can disrupt gene function (insertional mutagenesis) or activate nearby gene expression.
⭐ IS elements are fundamental units of transposition and often form parts of more complex transposons by flanking other genes (e.g., antibiotic resistance).
Transposon Types - Complex Jumpers
- Composite Transposons (e.g., Tn5, Tn9, Tn10)
- Structure: Central DNA (e.g., antibiotic resistance genes like $cat$, $tet$) flanked by two Insertion Sequences (IS elements).
- IS elements: Provide transposase & inverted repeats (IRs). Can be in direct or inverted orientation.
- Movement: Entire unit transposes.

- Non-Composite Transposons (e.g., Tn3, Tn21)
- Structure: No complete IS elements at ends; possess short terminal inverted repeats.
- Genes: Encode their own transposase, resolvase, and often accessory genes (e.g., $bla_{TEM}$ for ampicillin resistance).
- Mechanism: Often use replicative transposition (e.g., Tn3 involves a cointegrate intermediate).
⭐ Transposons are major contributors to the horizontal gene transfer and rapid spread of antibiotic resistance genes among bacteria, posing significant clinical challenges (e.g., MRSA, VRE).
Transposition Mechanisms - How They Hop
- Core Process: Transposons ("jumping genes") move using Transposase enzyme, which recognizes inverted repeats at the ends of the transposon.
- Two Major Pathways:
- Replicative Transposition ("Copy & Paste"):
- Transposon is duplicated during the process.
- Original copy is retained at the initial site; a new copy inserts elsewhere.
- Leads to an ↑ in the total number of transposons.
- Often involves a cointegrate intermediate, which is then resolved by Resolvase.
- E.g., $Tn3$ family.
- Non-Replicative/Conservative Transposition ("Cut & Paste"):
- Transposon is excised from the donor DNA site and inserted into a new target site.
- No net ↑ in transposon number (donor site is repaired, often imperfectly, or the DNA strand is lost).
- E.g., $Tn5$, $Tn10$.
- Replicative Transposition ("Copy & Paste"):
- Hallmark of Transposition: Creation of Target Site Duplications (TSDs) - short (3-12 bp) direct repeats of host DNA flanking the newly inserted transposon, resulting from staggered nicks made at the target site by transposase.

⭐ Exam Favourite: Replicative transposition results in an increase in the transposon's copy number within the genome, whereas conservative (non-replicative) transposition moves the transposon from one site to another without increasing its copy number.
Impact & Resistance - Genetic Shakers
- Genetic Impact:
- Insertional mutagenesis: Gene disruption/inactivation; can alter gene expression.
- Genomic rearrangements: Deletions, inversions, duplications (mediated by paired elements).
- Phase variation: Antigenic changes, e.g., Salmonella flagellar switching.
- Resistance Dissemination:
- Key mechanism for spreading antibiotic resistance genes (e.g., encoding β-lactamases, efflux pumps).
- Frequently found on plasmids (R-plasmids) or integrated into bacterial chromosomes.
- Composite transposons (Tn) often carry multiple resistance genes, conferring MDR.
⭐ Transposons are pivotal "genetic shakers," driving rapid evolution and dissemination of multi-drug resistance (MDR) among pathogenic bacteria.
 oka
oka
High‑Yield Points - ⚡ Biggest Takeaways
- Transposons ("jumping genes") are mobile DNA sequences that relocate within or between genomes.
- Insertion Sequences (IS) are simplest, encoding transposase and flanked by inverted repeats.
- Complex transposons (Tn) carry additional genes, notably for antibiotic resistance.
- Transposition is replicative (copy-paste) or non-replicative (cut-paste), mediated by transposase.
- They cause insertional mutations and are key in spreading antibiotic resistance genes.
- Target site duplications (short direct repeats) are formed flanking the newly inserted transposon.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

