CRISPR Fundamentals - Genome's Guardian
- CRISPR: Clustered Regularly Interspaced Short Palindromic Repeats.
- Natural Role: Adaptive immunity in bacteria/archaea against phages & plasmids.
- Key Locus Elements:
- Repeats: Short, palindromic DNA sequences.
- Spacers: Unique sequences derived from foreign DNA, interspersing repeats.
- Cas Genes: CRISPR-associated genes encoding proteins (e.g., nucleases).
- Leader Sequence: AT-rich region, directs CRISPR array transcription.

⭐ Spacers in the CRISPR array are derived from foreign DNA, acting as a molecular memory of past infections.
Mechanism of Action - DNA Scissors Dance
📌 AEI: Three stages:
- 1. Adaptation (Spacer Acquisition): Foreign DNA (protospacers) integrated as new spacers into CRISPR array.
- 2. Expression (crRNA Biogenesis): CRISPR array transcribed to pre-crRNA, processed to mature crRNAs. tracrRNA binds crRNA, forming a complex with Cas9 protein (or engineered as sgRNA with Cas9).
⭐ The PAM (Protospacer Adjacent Motif) is crucial for target DNA cleavage by Cas9. It's downstream of the target; its absence in the host CRISPR locus prevents self-targeting.
- 3. Interference (Target Cleavage): crRNA (within sgRNA for Cas9) guides Cas nuclease to complementary target DNA via base pairing. PAM sequence (e.g., $5'-NGG-3'$ for S. pyogenes Cas9) on target DNA is vital for Cas binding & cleavage (typically a Double-Strand Break/DSB by Cas9).

CRISPR System Types - The Cas Clan
- Class 1: Multi-subunit effector complexes.
- Class 2: Single effector protein (key for gene editing).
- Type II (Cas9): Targets DNA, causes Double-Strand Breaks (DSBs).
- Type V (Cas12a/Cpf1): DNA target, TTN PAM, staggered cuts, self-processes crRNA.
- Type VI (Cas13): Targets RNA.
Comparison: Cas9 vs. Cas12a
| Feature | Cas9 | Cas12a (Cpf1) |
|---|---|---|
| Nuclease | Cas9 | Cas12a |
| Target | DNA | DNA |
| PAM | NGG | TTN |
| Cut Type | Blunt DSB | Staggered DSB |
| gRNA | sgRNA (crRNA:tracrRNA) | crRNA (self-processed) |
Applications & Innovations - Gene Editing & Beyond
- Genome Editing:
- Gene knockout (NHEJ); Gene insertion/correction (HDR + donor template).
- Transcriptional Regulation: dCas9 (catalytically dead) with activators/repressors.
- Advanced Precision Editing:
- Base Editing: Deaminase-Cas9 fusion for point mutations (e.g., $C \rightarrow T$).
- Prime Editing: Versatile edits without extensive DSBs or donor DNA.
- Diagnostics:
- SHERLOCK (Cas13): Ultrasensitive RNA detection.
- DETECTR (Cas12): Rapid DNA detection.
- Therapeutics: Sickle cell disease, β-thalassemia, CAR-T cell therapy enhancement.
⭐ Base editors allow for precise single-nucleotide changes in DNA without requiring double-strand breaks, reducing the risk of indels associated with NHEJ.
oka
Limitations & Ethics - Hurdles & Headaches
- Technical Hurdles:
- Off-target effects: mutations at unintended sites.
- On-target large deletions/rearrangements.
- Mosaicism: mixed cell populations post-editing.
- Delivery challenges: efficient in vivo methods (viral, non-viral).
- Immunogenicity of Cas proteins.
- Ethical Headaches:
- Germline editing: heritable changes.
- Enhancement vs. therapy debate.
- Accessibility and equity.
- Informed consent complexities.
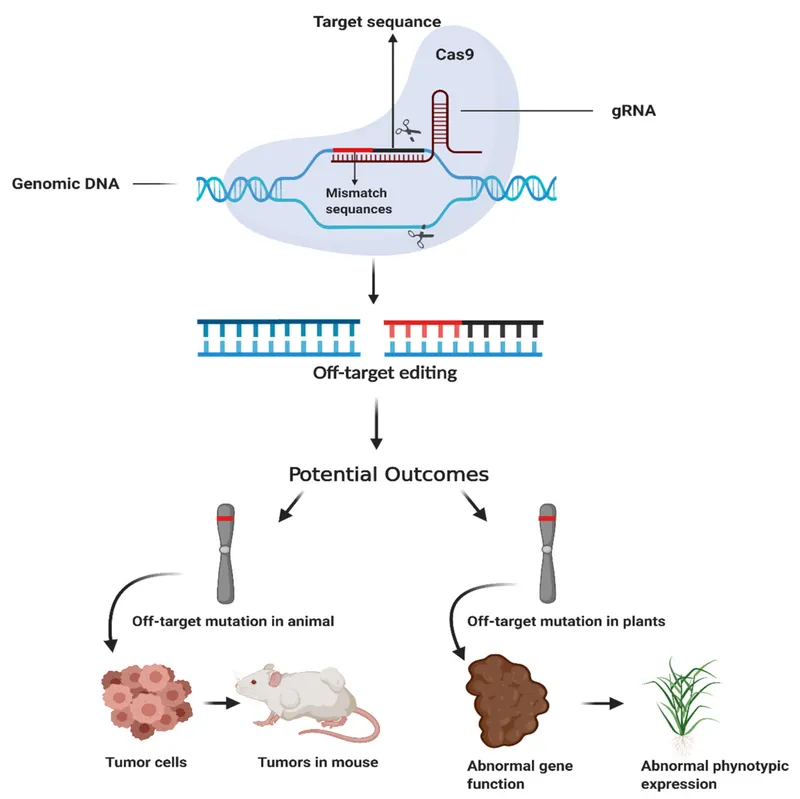
⭐ Off-target cleavage by Cas nucleases is a major safety concern for therapeutic applications, requiring careful guide RNA design and validation methods.
High‑Yield Points - ⚡ Biggest Takeaways
- CRISPR-Cas9: A prokaryotic adaptive immune system against phages/plasmids.
- Components: CRISPR arrays (repeats/spacers) & Cas proteins (e.g., Cas9 nuclease).
- Guide RNA (gRNA) directs Cas9 to complementary target DNA sequences.
- Cas9 induces double-strand breaks (DSBs) at the target, enabling gene editing.
- Applications: Gene knockout, knock-in, gene regulation, and diagnostics.
- PAM (Protospacer Adjacent Motif) sequence is vital for Cas9 target recognition and cleavage.
- Revolutionizing gene therapy, disease modeling, and antimicrobial research.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

