Bacterial Chromosome - The Main Blueprint
- Primary genetic element: typically single, circular, double-stranded DNA (dsDNA).
- Location: Cytoplasmic region called nucleoid; not membrane-bound.
- Ploidy: Usually haploid (one chromosome copy).
- Compaction:
- Supercoiling: Maintained by DNA gyrase (introduces negative supercoils) & topoisomerase IV (decatenation).
- Nucleoid-Associated Proteins (NAPs): e.g., HU, H-NS, IHF; histone-like proteins that help organize DNA.
- Size: Variable; E. coli genome is ~4.6 million base pairs (Mbp).
- Replication: Single origin of replication (oriC); proceeds bidirectionally.
- Genetic Content:
- High gene density; minimal non-coding DNA.
- No introns in protein-coding genes.
- Genes for related functions often grouped into operons.
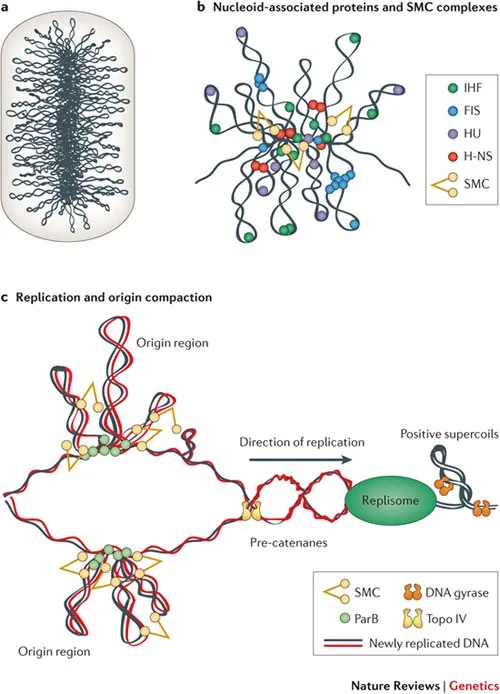
⭐ The enzyme DNA gyrase (a type II topoisomerase) is unique to bacteria and is a key target for quinolone antibiotics (e.g., ciprofloxacin).
Plasmids - Accessory Genetic Gear
-
Extrachromosomal, circular, dsDNA; self-replicating.
-
Smaller than chromosome; variable copy number (e.g., 1-100+).
-
Non-essential for viability; confer selective advantages (e.g., survival).
-
Key Types & Functions:
- R (Resistance) plasmids: Antibiotic resistance (e.g., β-lactamases, tetracycline resistance).
⭐ R-plasmids are key drivers of multidrug resistance (MDR) spread via horizontal gene transfer.
- F (Fertility) plasmids: Mediate conjugation (gene transfer via sex pili).
- Col plasmids: Produce bacteriocins (e.g., colicins), kill competing bacteria.
- Virulence plasmids: Encode toxins or adhesion factors (e.g., ETEC toxins, Bacillus anthracis pXO1/pXO2).
- Degradative plasmids: Metabolize unusual compounds (e.g., toluene).
- R (Resistance) plasmids: Antibiotic resistance (e.g., β-lactamases, tetracycline resistance).
-
Transfer: Horizontal (conjugation, transduction, transformation) & Vertical.
-
Curing: Spontaneous or induced plasmid loss. Incompatibility: Some plasmids cannot coexist in the same cell line without selection pressure (e.g. if they belong to the same incompatibility group Inc.).
Mobile Genetic Elements - Genome Shufflers
- DNA segments ("jumping genes") that move within or between genomes, mediating genetic variation.
- Types:
- Insertion Sequences (IS): Simplest; code only for transposase enzyme, flanked by inverted repeats (IRs).
- Transposons (Tn): Larger; carry additional genes (e.g., antibiotic resistance) besides transposition genes.
- Composite transposons: Central resistance genes flanked by IS elements.
- Non-composite transposons: Lack IS elements; have their own transposition genes.
- Mechanisms:
- Replicative ("Copy & Paste"): Element is duplicated, one copy moves.
- Non-replicative ("Cut & Paste"): Element moves directly without duplication.
- Significance:
- Major cause of mutations and genome rearrangements.
- Dissemination of antibiotic resistance (e.g., via R-plasmids carrying transposons).
- Facilitate bacterial evolution and diversity.
⭐ Transposons are a primary mechanism for the horizontal transfer of antibiotic resistance genes in bacteria.
Operons & Gene Regulation - Smart Gene Switches
- Operon: Prokaryotic gene cluster, co-regulated by single promoter/operator.
- Components: Regulator gene, Promoter (P), Operator (O), Structural Genes (SGs).
- Regulation Types:
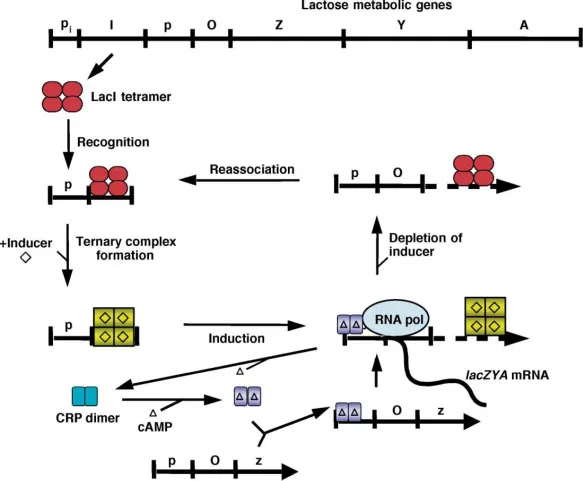
- Inducible (e.g., Lac Operon): Default OFF. Inducer (Allolactose) activates.
- Genes: lacZ, lacY, lacA.
- No lactose: Repressor binds Operator → OFF.
- Lactose: Allolactose binds repressor → inactive → ON.
⭐ Catabolite Repression: ↓Glucose → ↑cAMP. cAMP-CAP complex boosts lac operon if lactose present.
- Repressible (e.g., Trp Operon): Default ON. Co-repressor (Tryptophan) + Repressor deactivates.
- Genes: trpE,D,C,B,A (Trp synthesis).
- Low Trp: Repressor inactive → ON.
- High Trp: Trp (co-repressor) + Repressor → binds Operator → OFF.
- Inducible (e.g., Lac Operon): Default OFF. Inducer (Allolactose) activates.
- 📌 Mnemonic (Lac Operon): Lactose Absent, Genes Off (LAGO).
High-Yield Points - ⚡ Biggest Takeaways
- Single, circular chromosome is typical, located in the nucleoid (lacks nuclear membrane).
- Genomes are haploid, meaning one copy of each gene per cell.
- Plasmids: extrachromosomal circular DNA; often carry antibiotic resistance or virulence factors.
- Genes often organized into operons for coordinated expression (e.g., lac operon).
- Bacterial DNA: high gene density, minimal non-coding DNA, no introns.
- Mobile genetic elements (transposons, IS) contribute to genetic diversity and adaptation.
- Replication: bidirectional from a single origin of replication (oriC).
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

