Gene Expression Overview - Genes to Proteins
- Central Dogma (Bacteria): DNA $\xrightarrow{\text{Transcription}}$ RNA $\xrightarrow{\text{Translation}}$ Protein.
- Pathway: genetic info to functional proteins.
- Key Molecules & Structures:
- DNA: Double-stranded; stores genetic code.
- RNA: Single-stranded; key types:
- mRNA (messenger): Carries genetic code from DNA to ribosomes.
- tRNA (transfer): Delivers specific amino acids to ribosomes.
- rRNA (ribosomal): Major component of ribosomes; catalytic activity.
- Proteins: Functional molecules (e.g., enzymes, structural elements).
- Ribosomes: Cellular machinery for protein synthesis (translation).
- RNA Polymerase: Enzyme catalyzing transcription (DNA $\rightarrow$ RNA).
- Prokaryotic Hallmark: Transcription and translation are coupled (occur simultaneously in cytoplasm).
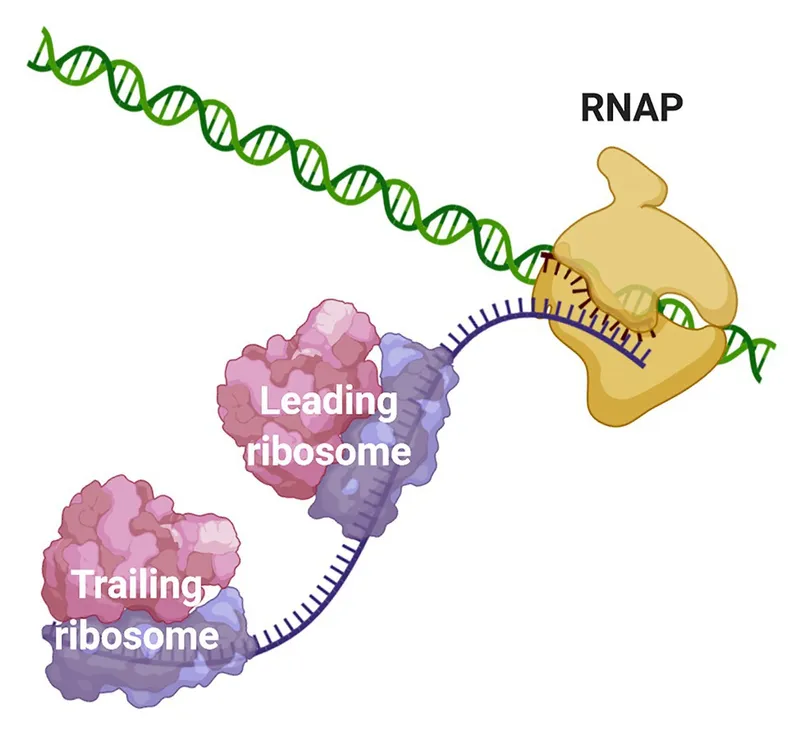
⭐ Bacterial mRNA: often polycistronic (one mRNA codes multiple proteins), short half-life; facilitates rapid environmental response.
Bacterial Transcription - Making the Message
Bacterial Transcription: DNA template → mRNA synthesis (5'→3').
- Enzyme: DNA-dependent RNA Polymerase (RNAP).
-
- Holoenzyme: Core enzyme (α₂ββ'ω) + Sigma (σ) factor.
-
- Sigma (σ) factor: Recognizes promoter, initiates transcription. Different σ factors for different gene sets (e.g., σ⁷⁰ for housekeeping).
-
- Promoter Elements (E. coli):
-
- Pribnow box (-10): TATAAT
-
- -35 sequence: TTGACA
- 📌 Mnemonic: "TTGACA" for -35, "TATAAT" for -10.
-
- Termination:
-
- Rho-dependent: Requires Rho protein (ATPase with helicase activity).
-
- Rho-independent (Intrinsic): GC-rich hairpin loop followed by a U-rich sequence in RNA transcript.
-
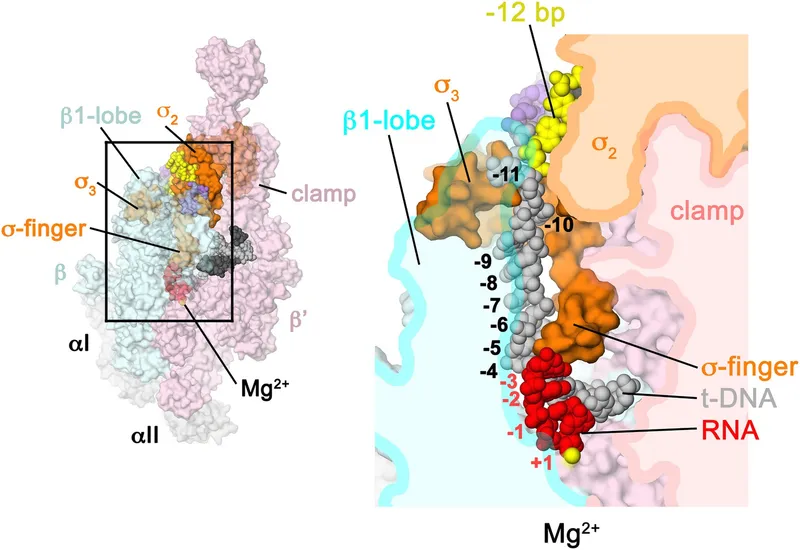
⭐ Rifampicin inhibits bacterial transcription by binding to the β-subunit of RNA polymerase, preventing initiation.
Bacterial Translation - Building the Blocks
- Ribosomes: 70S (30S + 50S). Site of protein synthesis.
- 30S subunit: Binds mRNA (via Shine-Dalgarno sequence upstream of AUG). Decodes mRNA.
- 50S subunit: Contains peptidyltransferase center (23S rRNA - a ribozyme); catalyzes peptide bond formation.
- Genetic Code: Triplet codons, non-overlapping, degenerate. Start: AUG (codes for N-formylmethionine - fMet in bacteria). Stop: UAA, UAG, UGA.
- tRNA: Adaptor molecule. Anticodon pairs with mRNA codon. Aminoacyl-tRNA synthetase ensures correct amino acid loading (charging).
- Process:
- 📌 Mnemonic for tRNA sites: A (Aminoacyl - Arrives) → P (Peptidyl - Peptide bond) → E (Exit).
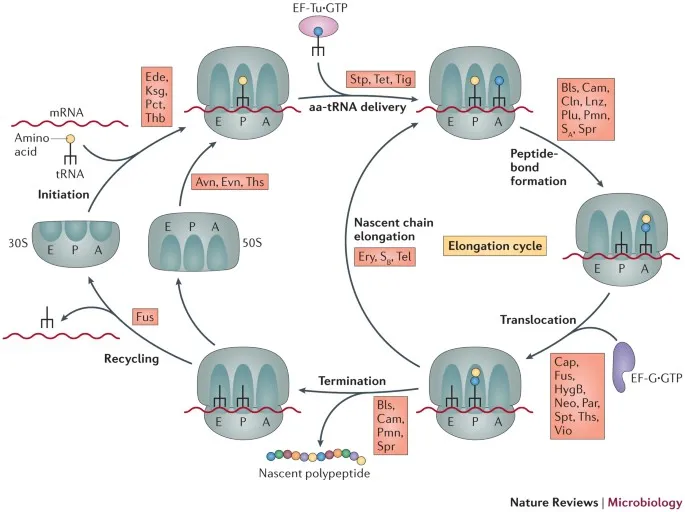
- 📌 Mnemonic for tRNA sites: A (Aminoacyl - Arrives) → P (Peptidyl - Peptide bond) → E (Exit).
⭐ Many antibiotics target bacterial translation: e.g., Aminoglycosides (Streptomycin) bind 30S causing misreading; Macrolides (Erythromycin) bind 50S blocking translocation.
Gene Regulation - Smart Switching
-
Bacteria use operons to regulate gene expression for adaptation.
-
Operon: Promoter, Operator, Structural genes. Regulator gene → repressor.
-
Lac Operon (Inducible - default OFF):
- Lactose metabolism. Inducer: Allolactose.
- No lactose: Repressor active, binds operator → transcription OFF.
- Lactose: Allolactose inactivates repressor → transcription ON.
- 📌 LACtose Is Needed (for INduction).
-
Trp Operon (Repressible - default ON):
- Tryptophan synthesis. Corepressor: Tryptophan.
- Trp low: Repressor inactive → transcription ON.
- Trp high: Trp activates repressor, binds operator → transcription OFF.
-
Catabolite Repression (Lac operon):
- Glucose preferred.
- ↑Glucose → ↓cAMP → CAP inactive → ↓Lac transcription.
- ↓Glucose → ↑cAMP → CAP active → ↑Lac transcription.
⭐ Glucose inhibits adenylate cyclase, ↓cAMP. cAMP is vital for CAP binding to enhance Lac operon transcription.

High‑Yield Points - ⚡ Biggest Takeaways
- Operons (e.g., lac, trp) are key for coordinated gene regulation in bacteria.
- Sigma factors direct RNA polymerase to specific promoters, initiating transcription.
- Coupled transcription-translation in the cytoplasm enables rapid cellular responses.
- Polycistronic mRNA encodes multiple proteins from one transcript, ensuring efficient pathway synthesis.
- Regulation primarily occurs at transcription initiation via inducers (e.g., allolactose) and repressors.
- Catabolite repression (glucose effect) prioritizes efficient energy sources, a global control mechanism.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

