Microbial Genetics

On this page
🧬 The Genetic Command Center: Bacterial DNA Architecture
Bacteria pack entire survival playbooks into chromosomes a thousand times smaller than ours, orchestrating gene expression through elegant operon switches, swapping resistance genes between species like encrypted messages, and even recording viral attacks in molecular memory banks. You'll discover how these microbial command centers control everything from nutrient scavenging to antibiotic resistance, why horizontal gene transfer makes bacteria evolution's most collaborative learners, and how clinicians now exploit bacterial genetics to diagnose infections and design precision antimicrobials that target genetic vulnerabilities.
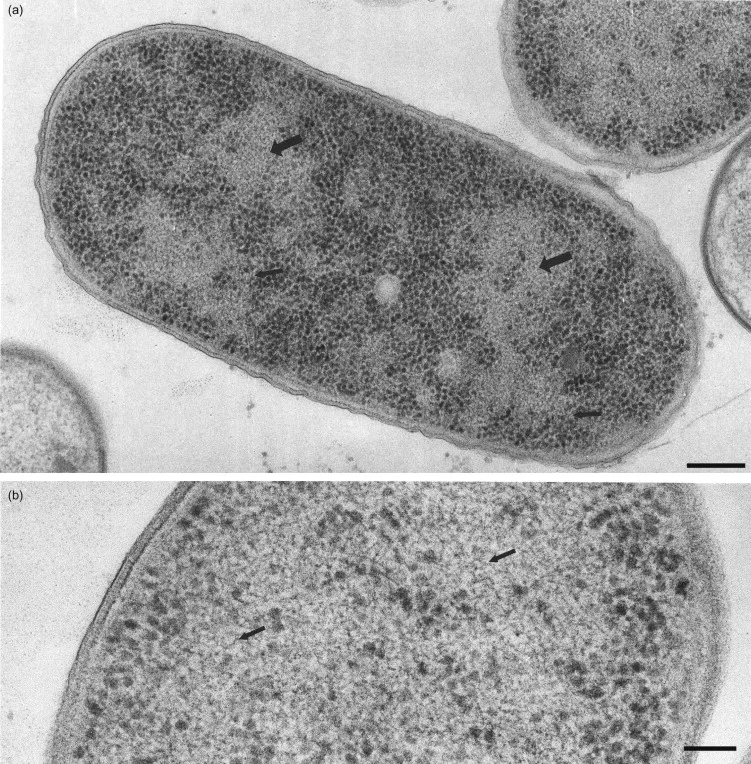
The bacterial genome represents a masterpiece of genetic efficiency, packing 4.6 million base pairs into a space 1,000 times smaller than a human cell nucleus. This remarkable organization enables rapid gene expression changes within 2-3 minutes of environmental shifts.
Core Genomic Architecture
-
Chromosomal Organization
- Single circular chromosome: 4.6 Mb average size
- Nucleoid region: 80% of cytoplasmic volume
- Supercoiling density: -0.06 (negative superhelical turns)
- DNA gyrase maintains optimal topology
- Topoisomerase I relieves excessive tension
- HU proteins organize nucleoid structure
-
Gene Density Patterns
- Coding sequences: 87% of total genome
- Intergenic regions: 13% (vs 98% in humans)
- Average gene length: 1,000 bp
- Overlapping genes: 15% of total
- Polycistronic transcripts: 60% of mRNAs
- Regulatory sequences: 100-200 bp upstream
📌 Remember: COMPACT - Circular chromosome, Overlapping genes, Minimal introns, Polycistronic mRNAs, Abundant operons, Coding density high, Tight regulation
| Feature | E. coli | S. aureus | M. tuberculosis | H. pylori | Clinical Significance |
|---|---|---|---|---|---|
| Genome Size | 4.6 Mb | 2.8 Mb | 4.4 Mb | 1.7 Mb | Metabolic complexity |
| Gene Number | 4,300 | 2,600 | 4,000 | 1,500 | Functional capacity |
| Coding % | 87% | 85% | 91% | 91% | Efficiency index |
| GC Content | 51% | 33% | 66% | 39% | Thermal stability |
| Plasmids | Variable | Common | Rare | None | Resistance potential |
Plasmid Architecture Systems
- Size Classifications
- Small plasmids: 1-10 kb (high copy number: 15-20 per cell)
- Medium plasmids: 10-100 kb (moderate copy: 5-10 per cell)
- Large plasmids: >100 kb (low copy: 1-3 per cell)
- Megaplasmids: >1 Mb (Rhizobium symbiotic functions)
- Chromids: Essential plasmids (500-1000 kb)
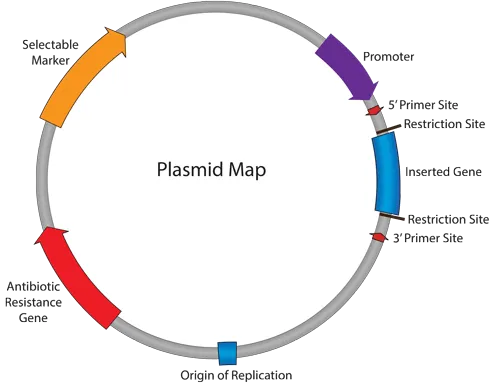
💡 Master This: Plasmid copy number inversely correlates with size - small plasmids achieve 20-fold higher expression levels than chromosomal genes, explaining why resistance genes on small plasmids create super-resistant phenotypes.
The sophisticated organization of bacterial genomes enables 10-fold faster protein synthesis compared to eukaryotes, with coupled transcription-translation occurring simultaneously. This efficiency becomes critical during stress responses, where survival depends on rapid metabolic reprogramming within minutes.
Connect this genomic foundation through operon organization to understand how bacteria coordinate hundreds of genes in synchronized responses to clinical interventions.
🧬 The Genetic Command Center: Bacterial DNA Architecture
⚙️ The Operon Engine: Coordinated Gene Control Networks
The Lac Operon: Metabolic Switch Mastery
The lactose operon exemplifies bacterial regulatory sophistication, integrating dual control mechanisms that respond to both substrate availability and glucose preference with mathematical precision.
-
Negative Control Architecture
- LacI repressor: 360 amino acids, tetrameric structure
- Operator binding: Kd = 10⁻¹¹ M (extremely tight)
- Allolactose induction: 1000-fold affinity reduction
- Inducer binding: 2 minutes for full derepression
- Basal transcription: 2% of maximum rate
- Full induction: 50-fold increase in mRNA
-
Positive Control Integration
- CAP-cAMP complex: glucose-sensitive activator
- Promoter binding: -61 to -72 position upstream
- Transcription enhancement: 20-fold increase
- cAMP levels: 10 μM (glucose absent) vs 1 μM (glucose present)
- CAP binding: Kd = 10⁻⁸ M with cAMP
- Synergistic activation: 1000-fold total range
📌 Remember: CAMP - CAP binding, Allolactose induction, Maximum when glucose absent, Positive and negative control
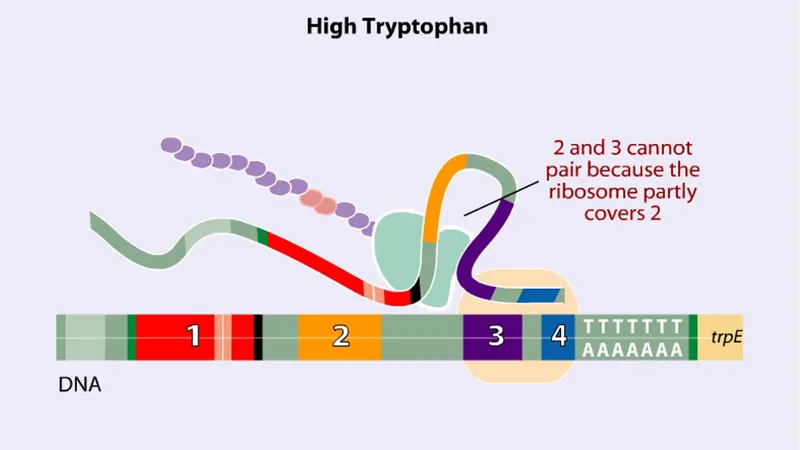
The Trp Operon: Attenuation Sophistication
The tryptophan operon demonstrates RNA-based regulation through attenuation, achieving fine-tuned control beyond simple repressor mechanisms.
- Attenuation Mechanism
- Leader sequence: 162 nucleotides with 4 stem-loop regions
- Tryptophan codons: 2 consecutive in leader peptide
- Termination efficiency: 90% (high tryptophan) vs 10% (low tryptophan)
- Ribosome stalling: 5-10 seconds at trp codons
- RNA polymerase: 30 nucleotides/second transcription rate
- Antitermination: 2:3 stem-loop formation
| Operon Type | Regulation | Response Time | Fold Change | Clinical Example |
|---|---|---|---|---|
| Catabolic (lac) | Inducible | 2-5 min | 50-1000x | Antibiotic degradation |
| Anabolic (trp) | Repressible | 1-3 min | 10-70x | Amino acid synthesis |
| Stress (heat shock) | Sigma factor | 30 sec | 5-20x | Fever response |
| Virulence (agr) | Quorum sensing | 10-30 min | 100-500x | Biofilm formation |
| SOS (recA) | DNA damage | 5-15 min | 50-200x | Mutation induction |
Regulatory Network Integration
- Global Regulators
- CRP-cAMP: Controls >100 operons (catabolite repression)
- FNR: >200 operons (anaerobic respiration)
- RpoS: >500 genes (stationary phase/stress)
- Sigma factor competition: 6 different sigma factors
- Promoter recognition: -10 and -35 consensus sequences
- Transcription initiation: 1-2 seconds per event
💡 Master This: Global regulators create regulatory hierarchies - CRP-cAMP controls 22% of E. coli genes, explaining why glucose availability affects antibiotic susceptibility across multiple resistance mechanisms simultaneously.
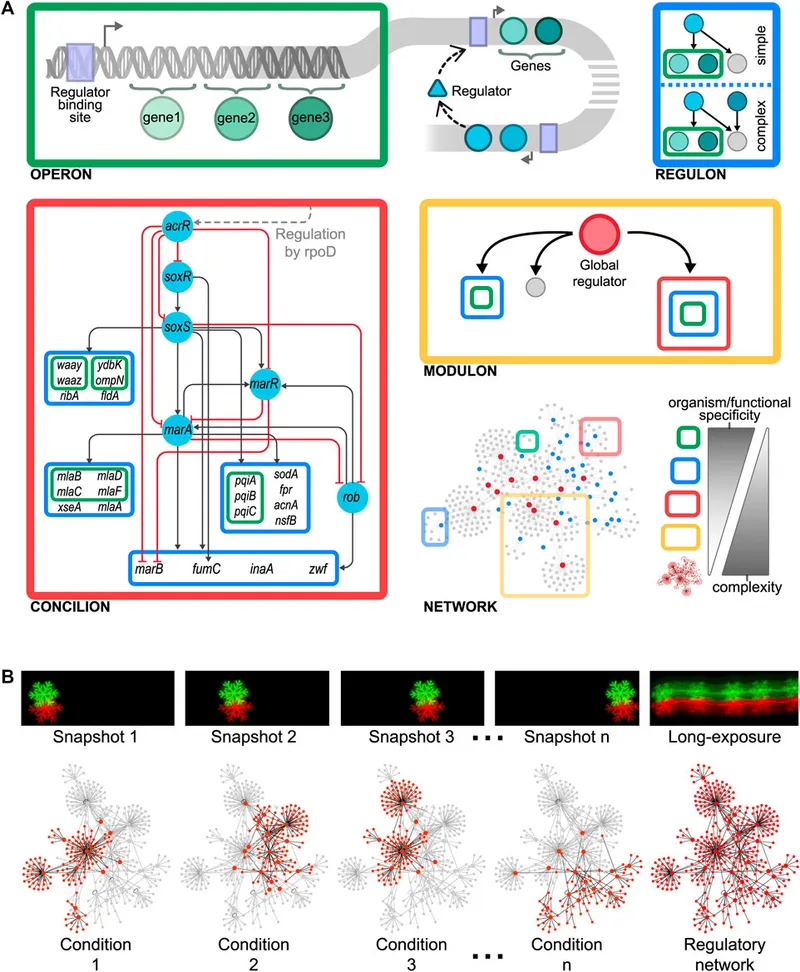
The operon system enables bacteria to coordinate metabolic responses with surgical precision, activating related pathways while silencing competitors. This regulatory sophistication allows rapid adaptation to antibiotic pressure within single generation times.
Connect operon coordination through mobile genetic elements to understand how resistance genes spread between regulatory contexts and create unpredictable phenotypes.
⚙️ The Operon Engine: Coordinated Gene Control Networks
🚀 Mobile Genetic Commandos: Horizontal Gene Transfer Networks
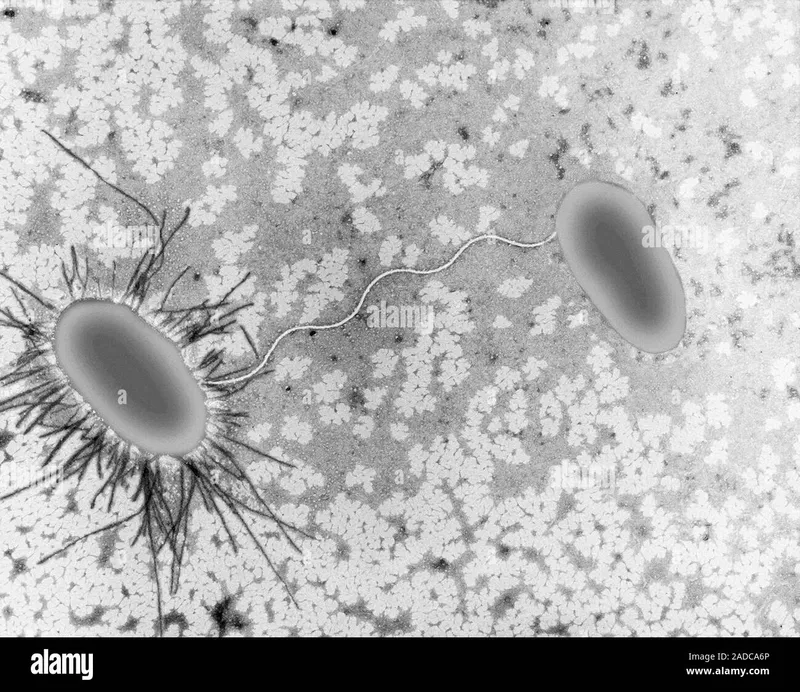
Conjugation: The Bacterial Mating Network
Conjugation represents the most sophisticated gene transfer mechanism, capable of moving megabase-sized DNA segments between distantly related bacterial species with remarkable efficiency.
-
Conjugative Machinery
- Type IV secretion system: 12-15 proteins forming transfer apparatus
- Pilus assembly: PilA protein polymerization (1000+ subunits)
- Transfer rate: 25-50 kb/minute through pilus channel
- Pilus length: 1-20 μm (up to 10x cell length)
- Contact time: 10-30 minutes for complete transfer
- Transfer efficiency: 10⁻² to 10⁻⁴ per donor cell
-
Plasmid Transfer Dynamics
- Rolling circle replication: 5' to 3' single-strand synthesis
- Relaxase enzyme: oriT recognition and nicking
- Complementary strand synthesis: simultaneous in recipient
- Transfer origin (oriT): 200-300 bp specific sequence
- Relaxosome complex: 8-12 proteins at transfer origin
- DNA processing: 3-5 kb segments transferred sequentially
📌 Remember: PILUS - Plasmid mobilization, Integration machinery, Large DNA transfer, Unidirectional movement, Single-strand synthesis
Transduction: Viral Gene Taxi Service
Bacteriophages serve as genetic vectors, transferring host DNA between bacterial cells with species specificity and high fidelity.
-
Generalized Transduction
- Packaging errors: 1-2% of phage particles contain host DNA
- DNA fragment size: 1-2% of bacterial genome (50-100 kb)
- Transfer frequency: 10⁻³ to 10⁻⁵ per phage particle
- Random packaging: any chromosomal region transferable
- Recipient integration: homologous recombination required
- Transductant frequency: 0.1-1% of infected cells
-
Specialized Transduction
- Prophage excision errors: imprecise excision events
- Adjacent gene transfer: 5-10 kb flanking prophage
- High frequency: 10⁻¹ to 10⁻² for specific loci
- Lambda phage: gal and bio operons (E. coli)
- P1 phage: any chromosomal region (large packaging capacity)
- Mu phage: transposition-mediated transfer
| Transfer Method | DNA Size Range | Species Barrier | Efficiency | Clinical Impact |
|---|---|---|---|---|
| Conjugation | 1 kb - 2 Mb | Low | 10⁻² - 10⁻⁴ | MDR plasmid spread |
| Transduction | 1-100 kb | High | 10⁻³ - 10⁻⁵ | Virulence factor transfer |
| Transformation | 0.1-50 kb | Medium | 10⁻⁴ - 10⁻⁶ | Competence-dependent |
| Transposition | 0.5-40 kb | None | 10⁻⁵ - 10⁻⁷ | Resistance gene mobility |
Transformation: Competence-Mediated Uptake
Natural transformation allows bacteria to acquire naked DNA from the environment, providing genetic flexibility during stress conditions.
- Competence Development
- Natural competence: species-specific (Streptococcus, Haemophilus, Neisseria)
- Competence factors: 10-15 proteins for DNA uptake
- Timing: late exponential to early stationary phase
- DNA binding proteins: ComEA (DNA receptor)
- DNA uptake machinery: ComEC (channel protein)
- DNA processing: DprA and RecA (integration)
💡 Master This: Competent bacteria preferentially uptake homologous DNA through species-specific sequences, explaining why Streptococcus pneumoniae readily acquires penicillin resistance from related streptococci but rarely from gram-negative sources.
The horizontal gene transfer network creates genetic highways between bacterial species, enabling resistance dissemination across taxonomic boundaries. This mobility explains why novel resistance combinations emerge rapidly and why infection control must consider entire microbial communities.
Connect mobile genetic elements through CRISPR-Cas systems to understand how bacteria defend against foreign DNA while selectively acquiring beneficial genes.
🚀 Mobile Genetic Commandos: Horizontal Gene Transfer Networks
🛡️ CRISPR-Cas Fortress: Adaptive Immunity Architecture
CRISPR Array Architecture
The CRISPR array functions as a genetic memory bank, storing spacer sequences derived from foreign DNA encounters with chronological organization.
-
Structural Components
- Spacer sequences: 20-50 nucleotides (foreign DNA fragments)
- Repeat sequences: 21-47 nucleotides (palindromic, conserved)
- Leader sequence: 100-500 bp (promoter and processing signals)
- Array expansion: unidirectional from leader end
- Spacer acquisition: 1-5% efficiency per infection
- Memory capacity: >200 spacers in some systems
-
Cas Protein Classifications
- Class 1 systems: multi-protein effector complexes (Types I, III, IV)
- Class 2 systems: single-protein effectors (Types II, V, VI)
- Cas9 (Type II): 1368 amino acids, dual nuclease domains
- HNH domain: target strand cleavage
- RuvC domain: non-target strand cleavage
- PAM recognition: NGG sequence requirement
📌 Remember: SPACER - Sequence-specific memory, PAM requirement, Adaptive immunity, Chronological order, Expansion from leader, Repeat-separated
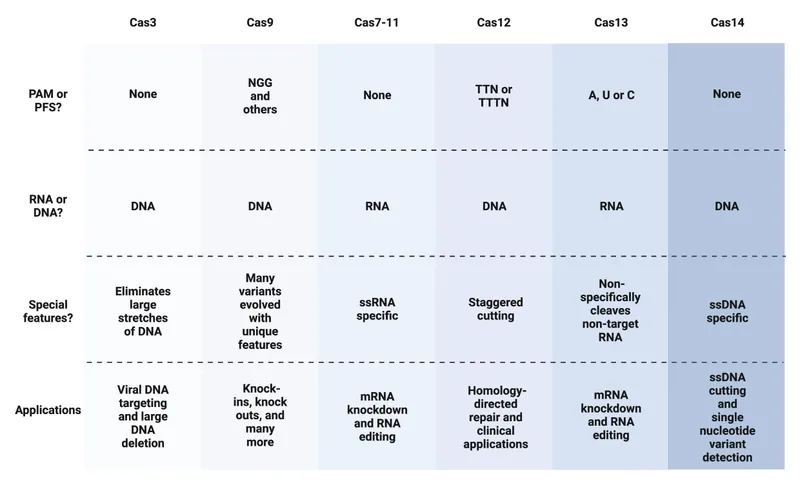
Type-Specific Mechanisms
Different CRISPR-Cas types employ distinct strategies for target recognition and DNA cleavage, each optimized for specific threats.
-
Type I Systems (Cascade Complex)
- Cas3 nuclease: 3' to 5' exonuclease and helicase activity
- Target degradation: progressive from PAM site
- Priming: enhanced spacer acquisition near degraded DNA
- Cascade binding: R-loop formation at target
- Cas3 recruitment: ATP-dependent translocation
- DNA degradation: >1000 bp from initial cut
-
Type II Systems (Cas9)
- Guide RNA: crRNA + tracrRNA hybrid (or single guide RNA)
- Cleavage pattern: blunt-ended double-strand breaks
- PAM requirement: 5'-NGG-3' (Streptococcus pyogenes)
- Target binding: 20 bp complementarity required
- Conformational change: PAM-induced Cas9 activation
- Cleavage timing: 3 bp upstream of PAM sequence
| CRISPR Type | Effector | Target | Cleavage Pattern | PAM Sequence | Clinical Relevance |
|---|---|---|---|---|---|
| Type I | Cascade + Cas3 | DNA | Progressive degradation | 5'-CTT-3' | Phage resistance |
| Type II | Cas9 | DNA | Blunt DSB | 5'-NGG-3' | Gene editing tool |
| Type III | Csm/Cmr | RNA/DNA | Sequence-specific | None | Transcription control |
| Type V | Cas12 | DNA | Staggered DSB | 5'-TTTV-3' | Multiplex editing |
| Type VI | Cas13 | RNA | RNA cleavage | None | RNA targeting |
Immune Evasion and Adaptation
CRISPR-Cas systems engage in evolutionary arms races with mobile genetic elements, driving rapid adaptation on both sides.
- Phage Counter-Strategies
- Anti-CRISPR proteins: >50 families identified
- PAM mutations: single nucleotide changes disable targeting
- Spacer mutations: 1-2 bp changes escape recognition
- AcrIIA4 (anti-Cas9): binds guide RNA and blocks cleavage
- DNA methylation: protects PAM sequences from recognition
- Homology avoidance: reduced similarity to host sequences
💡 Master This: CRISPR-Cas systems create selection pressure for anti-CRISPR evolution, explaining why therapeutic phages must be regularly updated and why phage cocktails show superior efficacy compared to single phage treatments.
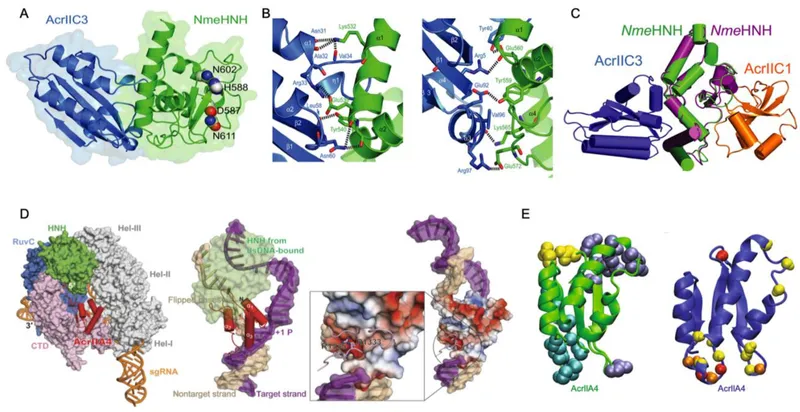
The CRISPR-Cas fortress demonstrates bacterial adaptive immunity sophistication, maintaining genetic memory while balancing foreign DNA defense with beneficial gene acquisition. This system's precision enables targeted gene editing applications while revealing natural mechanisms of genetic conflict.
Connect CRISPR defense systems through stress response networks to understand how bacteria coordinate immune responses with metabolic adaptation during clinical interventions.
🛡️ CRISPR-Cas Fortress: Adaptive Immunity Architecture
⚡ Stress Response Command: Survival Network Coordination
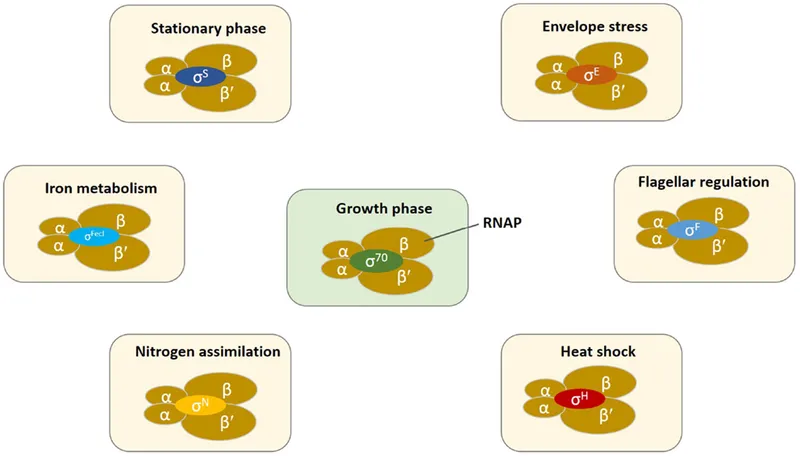
The RpoS General Stress Response
RpoS (σ³⁸) serves as the master regulator of bacterial stress adaptation, controlling >500 genes involved in survival and resistance mechanisms.
-
RpoS Regulation Hierarchy
- Transcriptional control: rpoS promoter responds to multiple signals
- Post-transcriptional: small RNAs (DsrA, RprA) stabilize rpoS mRNA
- Post-translational: RssB adaptor targets RpoS for ClpXP degradation
- Stress conditions: 10-fold increase in RpoS levels
- Half-life regulation: 1.4 minutes (exponential) vs 20 minutes (stationary)
- Activity control: anti-adaptor proteins (IraD, IraM, IraP)
-
RpoS-Controlled Processes
- Osmotic stress: trehalose synthesis, potassium uptake
- Oxidative stress: catalase, superoxide dismutase
- Acid resistance: glutamate decarboxylase, arginine deaminase
- Stationary phase survival: 100-fold increase
- Cross-protection: multiple stress tolerance
- Biofilm formation: curli fibers, cellulose production
📌 Remember: STRESS - Sigma 38 master, Transcriptional control, RNA regulation, Environmental sensing, Survival genes, Stationary phase
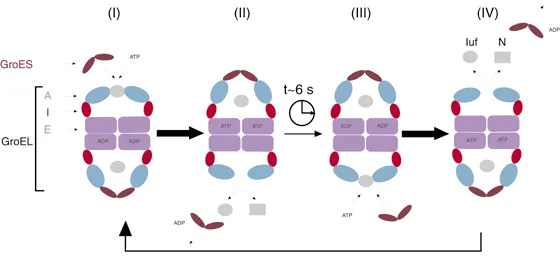
Heat Shock Response Networks
The heat shock response demonstrates rapid stress detection and protein quality control, essential for survival during fever and thermal stress.
- Sigma-32 (RpoH) System
- Temperature sensing: RNA thermometer in rpoH 5' UTR
- Protein targets: chaperones (GroEL, DnaK) and proteases (Lon, ClpP)
- Response kinetics: 30 seconds for initial induction
- GroEL/GroES: protein folding machinery (14-mer and 7-mer rings)
- DnaK/DnaJ/GrpE: co-chaperone system for nascent proteins
- Heat shock threshold: >42°C triggers 10-fold induction
| Stress Type | Sigma Factor | Response Time | Gene Number | Clinical Relevance |
|---|---|---|---|---|
| Heat Shock | σ³² (RpoH) | 30 seconds | ~30 | Fever survival |
| General Stress | σ³⁸ (RpoS) | 2-5 minutes | >500 | Antibiotic tolerance |
| SOS Response | σ⁷⁰ + LexA | 5-15 minutes | ~40 | DNA repair, mutagenesis |
| Stringent Response | σ⁷⁰ + (p)ppGpp | 1-3 minutes | >500 | Amino acid starvation |
| Cold Shock | σ⁷⁰ + CspA | 10-30 minutes | ~15 | Low temperature adaptation |
SOS Response: DNA Damage Control
The SOS response coordinates DNA repair and mutagenesis during severe DNA damage, often triggered by antibiotic treatment.
-
LexA-RecA Regulatory Circuit
- LexA repressor: controls >40 genes in SOS regulon
- RecA activation: single-strand DNA binding triggers coprotease activity
- LexA cleavage: auto-catalytic in presence of activated RecA
- DNA damage threshold: >50 double-strand breaks per genome
- RecA filament formation: cooperative binding to ssDNA
- Response magnitude: 10-200 fold gene induction
-
SOS-Induced Processes
- DNA repair: RecA, RecBCD, UvrABC systems
- Mutagenesis: UmuC/UmuD (DNA polymerase V) error-prone repair
- Cell division delay: SulA inhibits FtsZ polymerization
- Mutation rate increase: 100-1000 fold during SOS
- Survival benefit: 10-50% of cells survive lethal damage
- Resistance evolution: accelerated under SOS conditions
💡 Master This: SOS response creates hypermutable bacterial populations during antibiotic treatment - sub-lethal concentrations of fluoroquinolones induce SOS and accelerate resistance evolution by 10-100 fold.
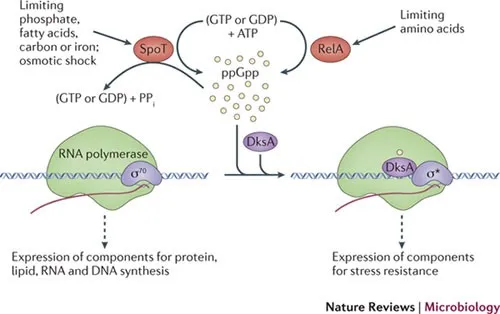
Stringent Response Integration
The stringent response coordinates metabolic shutdown and survival prioritization during nutrient limitation.
- (p)ppGpp Signaling
- RelA synthesis: uncharged tRNA binding to ribosomal A-site
- SpoT regulation: bifunctional enzyme (synthesis and hydrolysis)
- Global effects: >500 genes affected directly or indirectly
- rRNA synthesis: 90% reduction in ribosome production
- Amino acid biosynthesis: 5-20 fold upregulation
- Virulence factors: coordinate expression with growth phase
The stress response command center enables bacteria to survive clinical interventions through coordinated adaptation. These networks explain why combination therapies and rapid treatment protocols show superior efficacy compared to prolonged single-agent approaches.
Connect stress response coordination through resistance mechanisms to understand how environmental pressure drives genetic adaptation and clinical treatment failures.
⚡ Stress Response Command: Survival Network Coordination
🔬 Resistance Evolution Laboratory: Genetic Adaptation Mechanisms
Mutation-Driven Resistance Pathways
Spontaneous mutations provide the raw material for resistance evolution, with selection pressure determining which variants survive and proliferate.
-
Mutation Rate Dynamics
- Baseline rate: 10⁻⁹ to 10⁻¹⁰ per base pair per generation
- SOS-induced rate: 10⁻⁶ to 10⁻⁷ (100-1000 fold increase)
- Hypermutator strains: 10⁻⁵ to 10⁻⁶ (defective mismatch repair)
- DNA polymerase errors: 1 in 10⁵ nucleotides incorporated
- Proofreading efficiency: 99% error correction
- Mismatch repair: 99.9% of remaining errors corrected
-
Target-Specific Resistance Mutations
- β-lactam resistance: PBP mutations (>200 variants identified)
- Fluoroquinolone resistance: gyrA/parC mutations (>50 hotspots)
- Rifampin resistance: rpoB mutations (>30 resistance-conferring positions)
- Single-step resistance: 10⁻⁷ to 10⁻⁹ frequency
- Multi-step resistance: sequential accumulation over 5-10 generations
- Fitness costs: 5-20% growth rate reduction (compensatory mutations restore fitness)
📌 Remember: EVOLVE - Error-prone repair, Variation generation, Optimal selection, Large populations, Vertical transmission, Environmental pressure
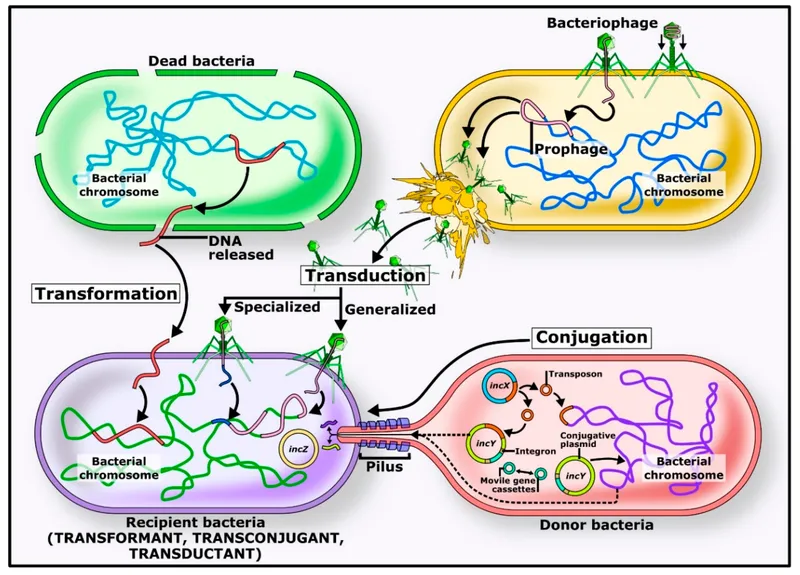
Horizontal Resistance Acquisition
Horizontal gene transfer accelerates resistance evolution by sharing pre-existing resistance mechanisms across bacterial species.
- Resistance Gene Mobilization
- Conjugative plasmids: >100 kb carrying multiple resistance genes
- Transposons: composite and complex structures mobilizing resistance
- Integrons: site-specific recombination systems capturing gene cassettes
- Class 1 integrons: >130 different resistance cassettes identified
- Transfer efficiency: 10⁻² to 10⁻⁴ per donor-recipient pair
- Multi-drug resistance: 5-15 genes per mobile element
| Resistance Mechanism | Mutation Rate | HGT Frequency | Time to Resistance | Stability |
|---|---|---|---|---|
| Target modification | 10⁻⁷ - 10⁻⁹ | N/A | 1-5 generations | High |
| Enzyme production | 10⁻⁹ - 10⁻¹¹ | 10⁻² - 10⁻⁴ | <1 generation | Variable |
| Efflux upregulation | 10⁻⁶ - 10⁻⁸ | 10⁻³ - 10⁻⁵ | 1-3 generations | Medium |
| Permeability reduction | 10⁻⁷ - 10⁻⁹ | Rare | 2-8 generations | High |
Population Dynamics and Selection
Bacterial populations employ sophisticated strategies to maximize survival under antibiotic pressure while maintaining genetic diversity.
-
Bet-Hedging Strategies
- Phenotypic heterogeneity: bistable gene expression creating resistant subpopulations
- Persister cells: 0.01-1% of population enters dormant state
- Competence development: stress-induced DNA uptake from environment
- Persistence duration: 6-24 hours of antibiotic tolerance
- Resuscitation rate: 10-90% depending on stress severity
- Genetic diversity: maintained through frequency-dependent selection
-
Adaptive Laboratory Evolution
- Serial passage: daily transfers under increasing drug concentrations
- Resistance development: 10-50 generations to high-level resistance
- Parallel evolution: similar mutations arise independently
- Predictable pathways: gyrA S83L in >80% of fluoroquinolone-resistant isolates
- Epistatic interactions: mutation order affects final resistance level
- Evolutionary constraints: limited pathways to high-level resistance
💡 Master This: Population bottlenecks during antibiotic treatment reduce genetic diversity and slow resistance evolution - high-dose, short-duration therapy exploits this principle by minimizing the time window for adaptation.
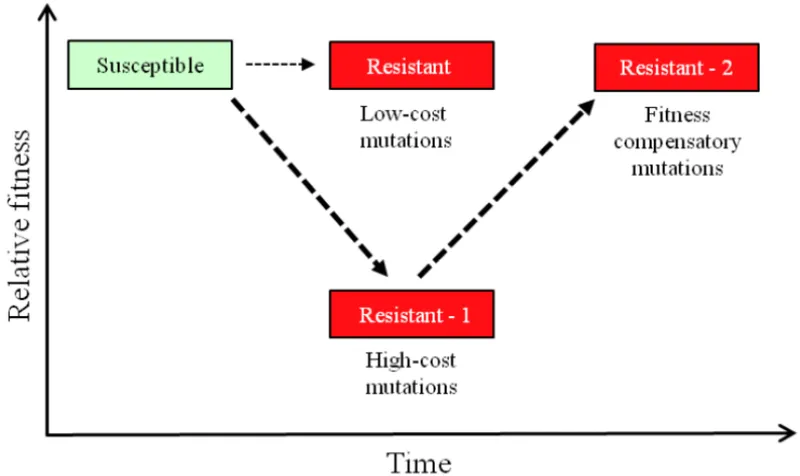
The resistance evolution laboratory demonstrates bacterial genetic plasticity and adaptive capacity under therapeutic pressure. Understanding these mechanisms enables rational design of evolution-resistant treatment strategies and combination therapies that minimize resistance development.
Connect resistance evolution through clinical applications to understand how laboratory insights translate into evidence-based therapeutic approaches and antimicrobial stewardship strategies.
🔬 Resistance Evolution Laboratory: Genetic Adaptation Mechanisms
🎯 Clinical Mastery Arsenal: Genetic Diagnostics and Therapeutic Targeting
Rapid Genetic Diagnostics
Modern molecular diagnostics leverage genetic signatures to provide actionable clinical information within 2-6 hours compared to 24-72 hours for traditional culture methods.
-
PCR-Based Detection Systems
- Multiplex PCR: 20-40 targets simultaneously (resistance genes + species identification)
- Real-time PCR: quantitative results with Ct values <30 for positive samples
- Digital PCR: absolute quantification without standard curves
- Sensitivity: 10-100 CFU/mL direct from clinical specimens
- Specificity: >99% for well-designed primer sets
- Turnaround time: 2-4 hours from sample to result
-
Next-Generation Sequencing Applications
- Whole genome sequencing: complete resistome and virulence profile
- Targeted amplicon sequencing: resistance gene variants and mutations
- Metagenomics: polymicrobial infections and microbiome analysis
- Cost reduction: <$100 per bacterial genome (2024 pricing)
- Turnaround time: 6-24 hours for clinical reporting
- Data output: 4-8 Gb per bacterial genome
📌 Remember: RAPID - Real-time detection, Amplification-based, Pathogen identification, Immediate results, Direct from specimen
Precision Antimicrobial Targeting
Genetic information enables mechanism-based therapy selection, optimizing drug choice, dosing, and combination strategies.
- Resistance Mechanism-Guided Therapy
- β-lactamase identification: specific inhibitor selection (clavulanate vs tazobactam vs avibactam)
- Efflux pump detection: pump inhibitor combinations or alternative drug classes
- Target modification: drug class switching based on specific mutations
- ESBL detection: carbapenem selection over cephalosporins
- Carbapenemase typing: combination therapy (meropenem + vaborbactam for KPC)
- MRSA identification: vancomycin or linezolid instead of β-lactams
| Resistance Gene | Mechanism | Affected Drugs | Optimal Alternative | Success Rate |
|---|---|---|---|---|
| blaCTX-M | ESBL | Cephalosporins | Carbapenems | >95% |
| blaKPC | Carbapenemase | Carbapenems | Ceftazidime-avibactam | 85-90% |
| mecA | PBP modification | β-lactams | Vancomycin/Linezolid | >90% |
| vanA | Target modification | Vancomycin | Linezolid/Daptomycin | 85-95% |
| aac(6')-Ib-cr | Aminoglycoside acetyltransferase | Aminoglycosides | Amikacin | 70-80% |
Therapeutic Optimization Strategies
- Pharmacokinetic-Pharmacodynamic Integration
- MIC prediction: genetic markers correlate with phenotypic resistance levels
- Dosing optimization: resistance mechanism determines optimal exposure targets
- Combination rationale: genetic profiles guide synergistic combinations
- β-lactam + β-lactamase inhibitor: ratio optimization based on enzyme kinetics
- Dual carbapenem therapy: genetic typing determines optimal combinations
- Efflux pump inhibitors: pump expression levels guide inhibitor dosing
💡 Master This: Genetic diagnostics enable precision antimicrobial therapy - patients receiving genotype-guided treatment show 25-40% higher cure rates and 50% fewer adverse events compared to empirical therapy approaches.
Clinical Implementation Framework
- Diagnostic Stewardship Integration
- Test selection: clinical syndrome + local epidemiology + patient factors
- Result interpretation: genetic findings + clinical context + treatment history
- Therapeutic adjustment: real-time modification based on genetic results
- Decision support systems: automated alerts for resistance-guided therapy
- Stewardship interventions: prospective review with genetic data
- Outcome monitoring: genetic markers predict treatment response
The clinical mastery arsenal transforms microbial genetics from academic knowledge into practical tools that improve patient outcomes. These approaches represent the future of infectious disease management, where genetic precision replaces empirical uncertainty in antimicrobial decision-making.
🎯 Clinical Mastery Arsenal: Genetic Diagnostics and Therapeutic Targeting
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app






















