General Microbiology

On this page
🔬 Microbial Universe: The Invisible Empire
You'll master the hidden world that shapes human health-from understanding how microbes build their cellular machinery and survive in diverse ecosystems, to recognizing pathogens through systematic clinical detective work. This lesson equips you to discriminate between organisms using precise diagnostic frameworks, deploy targeted antimicrobial strategies, and integrate ecological thinking into patient care. By connecting microbial structure to function and laboratory findings to treatment decisions, you'll develop the command center thinking that transforms you from memorizer to clinical microbiologist who sees patterns others miss.

📌 Remember: MICROBIAL - Morphology, Identification, Classification, Resistance, Organisms, Biochemistry, Infection, Antimicrobials, Laboratory methods form the core pillars of clinical microbiology mastery
The microbial world encompasses bacteria (>10,000 species), viruses (>5,000 types), fungi (>100,000 species), and parasites (>300 clinically significant). Understanding their fundamental characteristics enables rapid pathogen identification and targeted therapeutic interventions.
| Microorganism Type | Size Range | Cell Structure | Reproduction Time | Clinical Significance |
|---|---|---|---|---|
| Bacteria | 0.5-5 μm | Prokaryotic | 20 minutes-24 hours | Primary infectious agents |
| Viruses | 20-300 nm | Acellular | 6-48 hours | Intracellular parasites |
| Fungi | 2-200 μm | Eukaryotic | 1-7 days | Opportunistic pathogens |
| Parasites | 5 μm-10 m | Eukaryotic | Days to years | Complex life cycles |
| Prions | 2-5 nm | Protein only | Months to years | Neurodegenerative diseases |
- Bacterial Classification Hierarchy
- Domain: Bacteria (prokaryotic cellular organization)
- Phylum: Based on 16S rRNA sequencing (>97% similarity)
- Firmicutes: Gram-positive, low G+C content (<50%)
- Proteobacteria: Gram-negative, diverse metabolism
- Actinobacteria: High G+C content (>55%)
- Class: Metabolic and structural characteristics
- Order: Ecological and physiological groupings
- Family: Shared virulence factors and antibiotic resistance patterns
💡 Master This: Gram staining remains the first-line diagnostic tool because it correlates directly with cell wall structure, antibiotic susceptibility patterns, and initial empirical therapy selection in >90% of bacterial infections
The transition from morphological to molecular identification has revolutionized clinical microbiology, enabling species-level identification within 2-6 hours compared to traditional 24-72 hour culture methods.
🔬 Microbial Universe: The Invisible Empire
⚙️ Microbial Machinery: The Cellular Command Centers
📌 Remember: PEPTIDOGLYCAN - Protection, Elasticity, Permeability, Target for antibiotics, Identification marker, Determines shape, Osmotic regulation, Gram staining basis, Lysis prevention, Yield strength, Cross-linking, Antibiotic resistance, N-acetylmuramic acid backbone
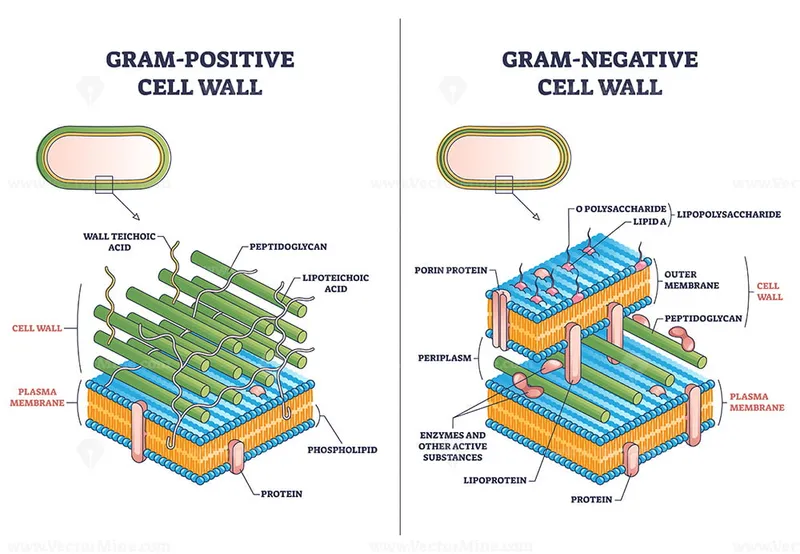
- Gram-Positive Cell Wall Architecture
- Peptidoglycan layer: 20-80 nm thick (90% of cell wall)
- Teichoic acids: Antigenic determinants and cation binding
- Wall teichoic acids: Structural integrity (40% of dry weight)
- Lipoteichoic acids: Membrane anchoring and adhesion
- Protein A (Staphylococcus): IgG binding (anti-phagocytic)
- M protein (Streptococcus): Complement resistance (virulence factor)
| Cell Wall Component | Gram-Positive | Gram-Negative | Clinical Significance |
|---|---|---|---|
| Peptidoglycan Thickness | 20-80 nm | 2-7 nm | β-lactam susceptibility |
| Outer Membrane | Absent | Present | Antibiotic barrier |
| Lipopolysaccharide | Absent | 10-15% | Endotoxin activity |
| Teichoic Acids | 40% | Absent | Antigenic variation |
| Porins | Absent | Present | Drug penetration |
- External Structures and Virulence
- Flagella: Motility apparatus (10-20 μm length, 20 nm diameter)
- H antigens: Serological typing (>170 types in Salmonella)
- Chemotaxis: Directed movement toward nutrients/away from toxins
- Pili (Fimbriae): Adhesion organelles (0.5-2 μm length)
- Type I pili: Mannose-sensitive adhesion (urinary tract)
- P pili: Kidney-specific binding (pyelonephritis)
- Sex pili: Conjugation and horizontal gene transfer
- Flagella: Motility apparatus (10-20 μm length, 20 nm diameter)
💡 Master This: Bacterial motility correlates with invasive potential - >80% of invasive E. coli strains are motile compared to <20% of commensal strains, making motility testing a virulence predictor
Understanding cellular architecture enables prediction of antibiotic susceptibility patterns and guides empirical therapy selection before culture results become available.
⚙️ Microbial Machinery: The Cellular Command Centers
🎯 Pathogen Recognition: The Clinical Detective Framework
📌 Remember: GRAM STAIN - General morphology, Rapid results, Antibiotic guidance, Membrane structure, Shape determination, Treatment selection, Arrangement patterns, Initial identification, Negative/positive classification
- Primary Identification Framework
- Gram Staining: Cell wall classification (>95% accuracy)
- Gram-positive: Purple/blue (crystal violet retention)
- Gram-negative: Pink/red (safranin counterstain)
- Gram-variable: Mixed staining (cell wall damage)
- Morphological Assessment: Shape and arrangement
- Cocci: Spherical (0.5-1.5 μm diameter)
- Bacilli: Rod-shaped (1-10 μm length)
- Spirilla: Spiral (helical morphology)
- Gram Staining: Cell wall classification (>95% accuracy)
| Organism Group | Gram Reaction | Morphology | Key Tests | Time to ID |
|---|---|---|---|---|
| Staphylococcus | Positive | Cocci clusters | Catalase + | 2-4 hours |
| Streptococcus | Positive | Cocci chains | Catalase - | 4-6 hours |
| Enterobacteriaceae | Negative | Bacilli | Oxidase - | 6-18 hours |
| Pseudomonas | Negative | Bacilli | Oxidase + | 12-24 hours |
| Mycobacterium | Acid-fast | Bacilli | AFB stain | 2-8 weeks |
- Biochemical Identification Cascade
- Catalase Test: H₂O₂ decomposition (bubble formation)
- Positive: Staphylococcus, Bacillus, Pseudomonas
- Negative: Streptococcus, Enterococcus, Clostridium
- Oxidase Test: Cytochrome c oxidase (color change)
- Positive: Pseudomonas, Neisseria, Vibrio (<10 seconds)
- Negative: Enterobacteriaceae (>99% specificity)
- Coagulase Test: Fibrinogen clotting (S. aureus identification)
- Positive: S. aureus (>95% sensitivity)
- Negative: Coagulase-negative Staphylococcus
- Catalase Test: H₂O₂ decomposition (bubble formation)
💡 Master This: The "3-minute rule" - Gram stain (1 minute), catalase (30 seconds), oxidase (30 seconds) provides preliminary identification sufficient for empirical antibiotic selection in >85% of bacterial infections
Modern molecular methods including MALDI-TOF MS achieve species-level identification within 5-15 minutes with >95% accuracy, revolutionizing clinical microbiology workflows and enabling rapid antimicrobial stewardship interventions.
🎯 Pathogen Recognition: The Clinical Detective Framework
🔍 Diagnostic Discrimination: The Precision Pathogen Matrix
📌 Remember: BIOCHEMICAL - Beta-hemolysis, Indole production, Oxidase activity, Catalase reaction, Hydrogen sulfide, Esculin hydrolysis, Motility testing, Iron utilization, Citrate utilization, Arginine hydrolysis, Lactose fermentation
- Gram-Positive Cocci Discrimination Matrix
- Staphylococcus vs. Streptococcus: Catalase test (100% discriminatory)
- Staphylococcus: Catalase positive (bubble formation)
- Streptococcus: Catalase negative (no bubbles)
- S. aureus vs. CoNS: Coagulase test (>95% accuracy)
- S. aureus: Coagulase positive (clot formation 4 hours)
- CoNS: Coagulase negative (no clotting)
- Streptococcus grouping: Hemolysis patterns (blood agar)
- α-hemolysis: Partial (greenish discoloration)
- β-hemolysis: Complete (clear zones)
- γ-hemolysis: None (no color change)
- Staphylococcus vs. Streptococcus: Catalase test (100% discriminatory)
| Organism | Catalase | Coagulase | Hemolysis | Bacitracin | Optochin |
|---|---|---|---|---|---|
| S. aureus | Positive | Positive | β | Resistant | Resistant |
| S. epidermidis | Positive | Negative | γ | Resistant | Resistant |
| S. pyogenes | Negative | Negative | β | Sensitive | Resistant |
| S. pneumoniae | Negative | Negative | α | Resistant | Sensitive |
| Enterococcus | Negative | Negative | α/β/γ | Resistant | Resistant |
- Gram-Negative Bacilli Discrimination
- Enterobacteriaceae vs. Non-fermenters: Oxidase test
- Enterobacteriaceae: Oxidase negative (>99% specificity)
- Pseudomonas: Oxidase positive (rapid color change)
- E. coli identification: IMViC pattern (classic biochemical series)
- Indole: Positive (tryptophan metabolism)
- Methyl red: Positive (mixed acid fermentation)
- Voges-Proskauer: Negative (butanediol pathway)
- Citrate: Negative (sole carbon source)
- Salmonella vs. Shigella: Key differentiating features
- Salmonella: H₂S positive, motile, citrate positive
- Shigella: H₂S negative, non-motile, citrate negative
- Enterobacteriaceae vs. Non-fermenters: Oxidase test
💡 Master This: Lactose fermentation on MacConkey agar provides immediate visual discrimination - pink colonies (lactose positive) suggest E. coli/Klebsiella while colorless colonies (lactose negative) suggest Salmonella/Shigella, guiding empirical therapy selection
Advanced molecular methods including 16S rRNA sequencing and MALDI-TOF mass spectrometry achieve species-level identification with >99% accuracy within minutes to hours, enabling rapid antimicrobial optimization.
🔍 Diagnostic Discrimination: The Precision Pathogen Matrix
⚖️ Treatment Precision: The Antimicrobial Strategy Engine
📌 Remember: ANTIBIOTIC - Allergy history, Nephrotoxicity, Tissue penetration, Interaction potential, Bacterial spectrum, Infection severity, Organ function, Toxicity profile, Immune status, Culture results

- Antimicrobial Mechanism Categories
- Cell Wall Synthesis Inhibitors: β-lactams (penicillins, cephalosporins)
- Target: Penicillin-binding proteins (PBPs)
- Mechanism: Peptidoglycan cross-linking inhibition
- Resistance: β-lactamase production (>50% S. aureus)
- Clinical use: First-line for susceptible Gram-positives
- Protein Synthesis Inhibitors: Aminoglycosides, macrolides
- Target: 30S/50S ribosomal subunits
- Mechanism: Translation disruption
- Resistance: Enzymatic modification (>30% Enterobacteriaceae)
- Cell Wall Synthesis Inhibitors: β-lactams (penicillins, cephalosporins)
| Antibiotic Class | Mechanism | Spectrum | Resistance Rate | Clinical Application |
|---|---|---|---|---|
| Penicillins | PBP inhibition | Gram-positive | >80% S. aureus | Streptococcal infections |
| Cephalosporins | PBP inhibition | Broad-spectrum | 15-30% E. coli | Surgical prophylaxis |
| Vancomycin | Cell wall synthesis | Gram-positive | <5% Enterococcus | MRSA infections |
| Fluoroquinolones | DNA gyrase | Broad-spectrum | >25% E. coli | UTI, respiratory |
| Aminoglycosides | 30S ribosome | Gram-negative | >40% Pseudomonas | Severe sepsis |
- Resistance Pattern Recognition
- MRSA Detection: Oxacillin/cefoxitin screening (>95% sensitivity)
- mecA gene: PBP2a production (β-lactam resistance)
- Treatment: Vancomycin, linezolid, daptomycin
- ESBL Producers: Cephalosporin resistance (>16 μg/mL)
- Mechanism: Extended-spectrum β-lactamases
- Treatment: Carbapenems (first-line therapy)
- Carbapenem Resistance: Meropenem MIC >2 μg/mL
- Mechanism: Carbapenemase production (KPC, NDM, OXA)
- Treatment: Colistin, tigecycline (salvage therapy)
- MRSA Detection: Oxacillin/cefoxitin screening (>95% sensitivity)
💡 Master This: Empirical therapy selection requires local antibiogram data - institutions with >20% MRSA prevalence need anti-MRSA coverage for serious Gram-positive infections, while <10% prevalence allows β-lactam monotherapy
Antimicrobial stewardship programs demonstrate 20-30% reduction in antibiotic consumption, 15-25% decrease in resistance rates, and $200,000-500,000 annual savings per hospital through optimized prescribing practices.
⚖️ Treatment Precision: The Antimicrobial Strategy Engine
🔗 Microbial Networks: The Ecosystem Integration Matrix
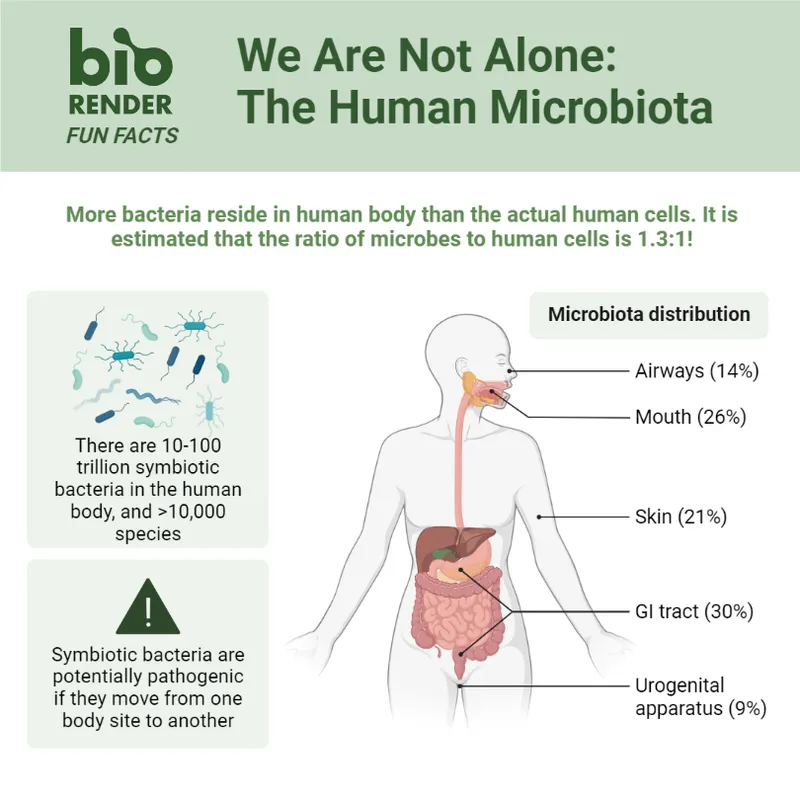
📌 Remember: MICROBIOME - Mucosal barriers, Immune modulation, Competitive exclusion, Resistance to colonization, Organ-specific communities, Biofilm formation, Infection susceptibility, Opportunistic pathogens, Metabolic functions, Ecological balance
- Normal Microbiota Distribution Patterns
- Skin Microbiome: >1,000 species (10⁶ CFU/cm²)
- Staphylococcus epidermidis: Dominant species (>50% abundance)
- Propionibacterium acnes: Sebaceous glands (anaerobic niches)
- Corynebacterium: Moist areas (axilla, groin)
- Gut Microbiome: >1,000 species (10¹¹-10¹² CFU/g)
- Bacteroidetes: >50% (fiber metabolism)
- Firmicutes: >30% (butyrate production)
- Proteobacteria: <10% (pathogen reservoir)
- Skin Microbiome: >1,000 species (10⁶ CFU/cm²)
| Body Site | Bacterial Load | Dominant Genera | Oxygen Level | Clinical Significance |
|---|---|---|---|---|
| Skin | 10⁶ CFU/cm² | Staphylococcus | Aerobic | Device infections |
| Oral cavity | 10⁸ CFU/mL | Streptococcus | Mixed | Endocarditis risk |
| Stomach | 10³ CFU/mL | Helicobacter | Microaerophilic | Ulcer disease |
| Small intestine | 10⁴ CFU/mL | Lactobacillus | Facultative | SIBO syndrome |
| Colon | 10¹² CFU/g | Bacteroides | Anaerobic | C. diff susceptibility |

- Pathogen-Host Interaction Networks
- Biofilm Communities: Multi-species aggregates (>1,000-fold antibiotic resistance)
- Extracellular matrix: Polysaccharides, proteins, DNA
- Quorum sensing: Cell density-dependent communication
- Clinical impact: >80% of device-associated infections
- Virulence Factor Networks: Coordinated pathogenicity
- Adhesins: Host cell attachment (tissue tropism)
- Toxins: Cellular damage (cytolytic, enterotoxic)
- Immune evasion: Complement resistance (capsule formation)
- Biofilm Communities: Multi-species aggregates (>1,000-fold antibiotic resistance)
💡 Master This: Polymicrobial infections show synergistic pathogenicity - anaerobic-aerobic combinations in abdominal infections demonstrate >50% higher mortality compared to monomicrobial infections, requiring broad-spectrum coverage
Understanding microbial networks enables precision medicine approaches that preserve beneficial microbiota while targeting pathogenic organisms, optimizing both therapeutic efficacy and microbiome stability.
🔗 Microbial Networks: The Ecosystem Integration Matrix
🎯 Clinical Mastery: The Microbiological Command Center
📌 Remember: CLINICAL MICRO - Culture interpretation, Lab result integration, Infection control, Normal flora knowledge, Identification methods, Contamination recognition, Antibiotic selection, Laboratory communication, Molecular diagnostics, Isolation procedures, Critical values, Resistance patterns, Outbreak investigation
- Essential Clinical Arsenal
- Gram Stain Mastery: <5 minutes to preliminary diagnosis
- Sensitivity: >95% for bacterial detection
- Specificity: >90% for organism classification
- Clinical impact: Immediate empirical therapy guidance
- Culture Interpretation: 24-72 hours to definitive results
- Quantitative cultures: >10⁵ CFU/mL (significant bacteriuria)
- Blood cultures: >1 organism (true bacteremia vs. contamination)
- Respiratory cultures: >10⁶ CFU/mL (pneumonia threshold)
- Gram Stain Mastery: <5 minutes to preliminary diagnosis
| Clinical Scenario | Key Organisms | Empirical Therapy | Diagnostic Timeline |
|---|---|---|---|
| UTI (uncomplicated) | E. coli (85%) | Nitrofurantoin | 1-2 days |
| Pneumonia (CAP) | S. pneumoniae (30%) | Amoxicillin | 2-3 days |
| Skin/soft tissue | S. aureus (40%) | Clindamycin | 1-2 days |
| Bacteremia | Mixed flora | Broad-spectrum | 2-5 days |
| Meningitis | S. pneumoniae (50%) | Ceftriaxone + Vancomycin | <6 hours |
- Rapid Diagnostic Integration
- Molecular Methods: PCR, MALDI-TOF (<2 hours results)
- Sensitivity: >95% for pathogen detection
- Specificity: >98% for species identification
- Cost-effectiveness: $50-200 per test vs. $500-2000 treatment delay
- Antigen Detection: Urinary antigens (<30 minutes)
- S. pneumoniae: >90% sensitivity in bacteremic pneumonia
- Legionella: >95% specificity for serogroup 1
- Molecular Methods: PCR, MALDI-TOF (<2 hours results)
💡 Master This: Critical value communication requires <1 hour notification for positive blood cultures, CSF organisms, or resistant pathogens - delays >2 hours correlate with increased mortality (>20% relative risk)
Clinical microbiology expertise enables evidence-based medicine through rapid pathogen identification, targeted antimicrobial therapy, and infection control optimization, directly improving patient outcomes and healthcare efficiency.
🎯 Clinical Mastery: The Microbiological Command Center
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app




















