Bacterial Identification Techniques - Stains & Scopes
- Gram Stain: 📌 CIAS (Crystal Violet→Iodine→Alcohol→Safranin).
- Gram +ve: Purple (e.g., Staphylococcus, Streptococcus). Thick peptidoglycan.
- Gram -ve: Pink/Red (e.g., E. coli). Thin peptidoglycan, outer membrane.
- Ziehl-Neelsen (ZN) Stain: For Acid-Fast Bacilli (AFB) like Mycobacterium tuberculosis (red rods); background blue.
- Special Stains:
- India Ink: Negative stain for capsules (e.g., Cryptococcus neoformans).
- Albert's Stain: C. diphtheriae metachromatic granules (green bacilli, blue-black granules).
- Microscopy Types:
- Bright-field: Routine stained preps.
- Dark-field: Spirochetes (e.g., Treponema).
- Phase Contrast: Unstained, live organisms.
- Fluorescence: Auramine-Rhodamine (TB), immunofluorescence.

⭐ The cell wall difference (peptidoglycan thickness and outer membrane presence) is the primary reason for Gram stain differentiation.
Bacterial Identification Techniques - Culture Club
- Culture Media Types:
- Basal: Nutrient agar (general growth)
- Enriched: Blood agar (BA), Chocolate agar (CA) (fastidious organisms) 📌 Blood & Chocolate for the demanding guests!
- Selective: MacConkey agar (Gram-negative), Lowenstein-Jensen (LJ) medium (M. tuberculosis)
- Differential: MacConkey (lactose fermenters pink), CLED agar (lactose fermenters yellow)
- Transport: Stuart's, Amies (preserve specimen viability)
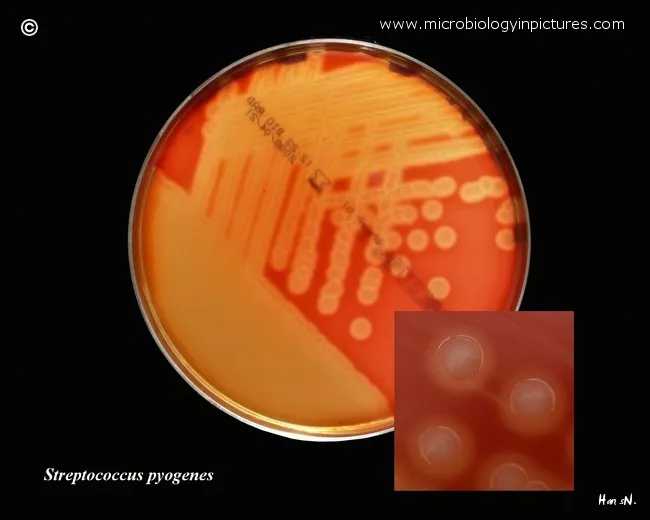
- Colony Characteristics: Size, shape, elevation, margin, color, opacity, consistency, hemolysis on BA (α, β, γ).
- Atmospheric Requirements:
- Aerobic: Requires O₂ (e.g., Pseudomonas)
- Anaerobic: O₂ is toxic (e.g., Clostridium)
- Microaerophilic: Low O₂ (e.g., Campylobacter at 5-10% O₂, 42°C)
- Capnophilic: High CO₂ (e.g., Neisseria, Haemophilus)
⭐ Chocolate agar is prepared by heating blood agar, lysing RBCs, releasing Factor X (hemin) & Factor V (NAD), vital for Haemophilus influenzae & Neisseria gonorrhoeae growth at 37°C.
Bacterial Identification Techniques - Bio-Chem Detectives
- Catalase: Detects enzyme breaking $H_2O_2$. $2H_2O_2 \rightarrow 2H_2O + O_2$.
- Oxidase: Identifies cytochrome c oxidase (e.g., Pseudomonas).
- Coagulase: Differentiates S. aureus (positive) from CoNS. Slide (bound), Tube (free).
- Urease: Detects urease enzyme (e.g., Proteus, H. pylori).
- Indole: Tests for tryptophanase enzyme.
- IMViC Reactions:
- Indole, Methyl Red (mixed acid), Voges-Proskauer (acetoin), Citrate (sole carbon source).
- 📌 E. coli: ++--; Klebsiella/Enterobacter: --++.
- TSI Agar (Triple Sugar Iron):
- Fermentation: Glucose (K/A), Lactose/Sucrose (A/A).
- H₂S production (black precipitate), Gas production.
 , A/A (lactose/sucrose fermenter), H₂S production)
, A/A (lactose/sucrose fermenter), H₂S production)
- Automated Systems: VITEK, API strips for rapid identification.
⭐ The oxidase test is crucial for differentiating Enterobacteriaceae (oxidase-negative, except Plesiomonas) from Pseudomonas species (oxidase-positive).
Bacterial Identification Techniques - Sero & Geno ID
- Immunological Methods (Serological ID)
- Agglutination: Detects Ag-Ab clumping.
- Slide/Tube: Serotyping (e.g., Salmonella, Shigella).
- Latex: Lancefield grouping (Streptococci), Cryptococcus.
- Precipitation: Ag-Ab reaction in gel/solution (e.g., VDRL).
- ELISA (Enzyme-Linked Immunosorbent Assay): Detects Ag or Ab (e.g., HIV).

- Immunofluorescence (IF): Ag detection using fluorescent Abs.
- Direct (DFA): e.g., Chlamydia trachomatis.
- Indirect (IFA): e.g., ANA.
- Agglutination: Detects Ag-Ab clumping.
- Molecular Methods (Genotypic ID)
- PCR (Polymerase Chain Reaction): Amplifies DNA/RNA.
- RT-PCR: RNA targets (e.g., RNA viruses).
- Multiplex PCR: Multiple targets simultaneously.
- 16S rRNA Gene Sequencing: Gold standard for bacterial phylogeny & ID.
- MALDI-TOF MS: Rapid ID via protein profiling.
- DNA Probes & RFLP/PFGE: Epidemiological typing.
- PCR (Polymerase Chain Reaction): Amplifies DNA/RNA.
⭐ MALDI-TOF MS has revolutionized clinical microbiology by providing rapid (minutes) and accurate identification of bacteria and fungi directly from colonies or even positive blood cultures, based on their unique ribosomal protein fingerprint.
- Comparison: Immunological vs. Molecular
- Immunological: Detect host response or microbial Ags; established, cost-effective for some.
- Molecular: Detect microbial nucleic acids; high sensitivity/specificity, ID non-culturables, rapid for some (MALDI-TOF, PCR).
High‑Yield Points - ⚡ Biggest Takeaways
- Gram stain is the crucial first step in bacterial differentiation.
- Ziehl-Neelsen stain identifies acid-fast bacilli like Mycobacterium tuberculosis.
- Culture on appropriate media is vital for isolation and preliminary ID.
- Biochemical tests (e.g., catalase, oxidase, IMViC) are key for species-level ID.
- Serological tests like agglutination identify specific antigens.
- Molecular methods like PCR offer rapid, sensitive, and specific identification.
- MALDI-TOF MS allows rapid, accurate identification based on protein profiles.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more