Introduction & Pre-Analytical Automation - Robo-Plates & Codes
- Why Automate? ↑Efficiency, ↑Accuracy, ↓Turnaround Time (TAT), ↓Manual errors, ↓Labor costs. Standardizes processes, improves traceability.
- Pre-Analytical Automation: Focuses on specimen processing before analysis.
- Codes (Barcoding): Crucial for positive patient ID. Uses 1D (linear) or 2D (e.g., QR codes) barcodes. Enables automated specimen reception, sorting, and tracking. Integrates with Laboratory Information Systems (LIS).
- Robo-Plates (Automated Inoculation): Systems like WASP (Walk-Away Specimen Processor) or Kiestra TLA/PLA automate media streaking. Ensures consistent, high-quality inoculation patterns, reducing variability.
⭐ Barcoding is paramount in pre-analytical automation, significantly minimizing patient identification and specimen handling errors, thereby enhancing patient safety.

Automated Culture & Rapid ID (MALDI-TOF) - Culture Bots & Speedy IDs
- Automated Culture Systems (e.g., Blood Cultures):
- Examples: BACTEC, BacT/ALERT, VersaTREK.
- Principle: Continuous monitoring for microbial growth (CO2 detection - colorimetric/fluorescent, or pressure changes).
- Benefits: Faster detection (often 24-72 hrs), reduced hands-on time, standardization, early alerts.
- MALDI-TOF MS (Matrix-Assisted Laser Desorption/Ionization Time-of-Flight):
- Rapid ID of bacteria, yeasts, fungi from colonies or processed positive blood cultures.
- Principle: Generates unique protein (mainly ribosomal) fingerprint.
- Process: Organism + matrix → laser ionization → ions separated by mass-to-charge ratio (m/z) in flight tube → spectrum compared to database.
- Advantages: ID in minutes, high accuracy, cost-effective per test (after initial instrument cost).
⭐ MALDI-TOF MS reduces identification time from 24-48 hours (conventional biochemical tests) to <1 hour post-culture.
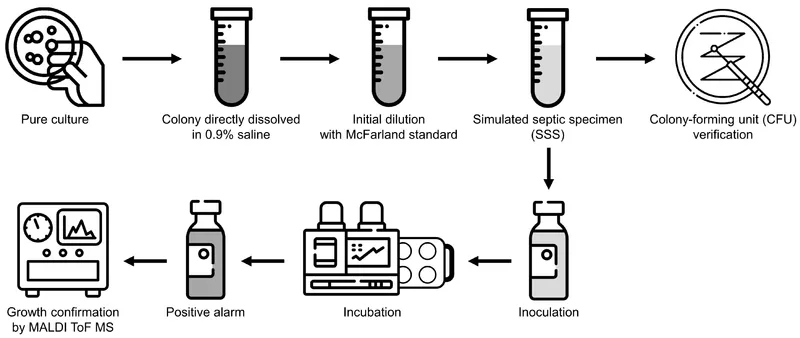
Automated ID & AST Systems - Smart Tests, Smart Bugs
- Automate microbial ID & antimicrobial susceptibility testing (AST).
- Principles:
- ID: Biochemical reactions (colorimetric, fluorogenic, turbidimetric), enzyme detection.
- AST: Automated broth microdilution (BMD) for MICs; turbidity/color change monitoring.
- Key Systems:
- VITEK (bioMérieux): Compact cards, biochemical tests.
- MicroScan WalkAway (Beckman Coulter): Microtiter panels.
- BD Phoenix: Broth-based, redox indicators.
- Advanced ID: MALDI-TOF MS (VITEK MS, Bruker Biotyper) for rapid proteomic fingerprinting.
- Benefits: ↑Speed (results 4-24 hrs), ↑accuracy, standardization, ↓turnaround time, LIS integration.
⭐ Many systems use expert rules to detect resistance mechanisms (e.g., ESBL, MRSA).
Post-Analytics, Molecular Automation & Trends - Data Flow & Crystal Ball
- Post-Analytics & Data Flow:
- LIS/LIMS: Central for data management, result integration, audit trails.
- Auto-validation: Rule-based algorithms for result release, flagging outliers, ↓errors.
- Critical value alerts: Automated, ensuring timely clinician notification.
- Data warehousing & Analytics: Supports epidemiology, AMR trend analysis, quality control.
- Molecular Automation Advances:
- Key Systems: GeneXpert (cartridge-based), BioFire FilmArray (syndromic), automated NA extraction & PCR setup.
- Benefits: ↑Throughput, ↓Turn-Around Time (TAT), ↑Reproducibility, ↓Manual labor.
- NGS Automation: Streamlining Whole Genome Sequencing for outbreak investigation & comprehensive AMR profiling.
- Future Outlook (Crystal Ball):
- AI/ML: For AMR prediction from genomic data, automated microscopy, outbreak detection.
- Total Lab Automation (TLA): Integrating molecular workflows into consolidated systems.
- Advanced Point-of-Care (POC) molecular tests: Decentralizing diagnostics.
- Enhanced data interoperability with Electronic Health Records (EHRs).
⭐ AI algorithms analyzing MALDI-TOF MS spectra or WGS data can predict antimicrobial resistance patterns faster than conventional AST.
 oka
oka
High‑Yield Points - ⚡ Biggest Takeaways
- Automation significantly reduces Turnaround Time (TAT) for microbial ID & AST.
- MALDI-TOF MS ensures rapid, accurate species-level identification from cultures.
- Automated blood culture systems (e.g., BacT/ALERT, BACTEC) enable early sepsis detection via continuous monitoring.
- Automated AST systems (e.g., VITEK, MicroScan) provide standardized and reliable susceptibility profiles.
- Automated molecular platforms allow swift detection of pathogens & key resistance genes.
- Total Lab Automation (TLA) integrates processes, enhancing efficiency and reducing errors.
- Core advantages: ↑accuracy, ↑reproducibility, ↓manual errors, improved safety.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more


