Diagnostic Microbiology

On this page
🔬 The Diagnostic Detective's Arsenal: Microbiology Mastery
Diagnostic microbiology transforms invisible threats into actionable intelligence, bridging the gap between a patient's symptoms and life-saving treatment. You'll master the complete diagnostic pathway-from precise specimen collection that preserves pathogen viability, through culture techniques that coax organisms into visibility, to rapid tests that deliver answers in minutes rather than days. You'll decode antimicrobial susceptibility patterns that guide therapy, harness molecular methods that identify pathogens by their genetic fingerprints, and integrate these tools into clinical decision-making that changes outcomes. This is detective work at the microscopic scale, where your technical precision and interpretive skill directly determine whether antibiotics hit their mark.

📌 Remember: CLSI - Clinical Laboratory Standards Institute guidelines govern 95% of diagnostic microbiology protocols worldwide
The diagnostic journey begins with specimen collection and progresses through multiple analytical phases, each requiring specific technical expertise and quality control measures. Modern laboratories integrate traditional culture methods with rapid molecular diagnostics, creating comprehensive pathogen detection systems.
- Pre-analytical Phase (40% of errors occur here)
- Specimen collection timing and technique
- Transport conditions and media selection
- Temperature maintenance: 2-8°C for most specimens
- Transport time limits: <2 hours for fastidious organisms
- Analytical Phase (30% of workflow time)
- Primary culture setup and incubation
- Microscopic examination and staining
- Gram stain sensitivity: 60-90% depending on specimen type
- Direct antigen detection: 85-95% specificity for targeted pathogens
- Post-analytical Phase (Critical interpretation)
- Result validation and clinical correlation
- Antimicrobial susceptibility reporting
| Diagnostic Method | Time to Result | Sensitivity | Specificity | Cost Factor | Clinical Application |
|---|---|---|---|---|---|
| Culture Methods | 24-72 hours | 90-95% | >99% | Low | Gold standard identification |
| Rapid Antigen | 15-30 minutes | 70-85% | 95-99% | Medium | Point-of-care screening |
| Molecular PCR | 2-6 hours | 95-99% | 98-100% | High | Rapid definitive diagnosis |
| MALDI-TOF MS | 5-15 minutes | 85-95% | >95% | Medium | Rapid organism identification |
| Automated Systems | 4-18 hours | 90-98% | 95-99% | High | High-throughput processing |
💡 Master This: Every diagnostic decision depends on understanding the analytical sensitivity (10²-10⁴ CFU/mL detection limits) versus clinical sensitivity (correlation with disease presence) for each testing methodology
The integration of traditional and molecular methods creates diagnostic algorithms optimized for specific clinical scenarios. Understanding these workflows enables clinicians to select appropriate testing strategies and interpret results within proper clinical context.
🔬 The Diagnostic Detective's Arsenal: Microbiology Mastery
🎯 Specimen Collection Mastery: The Foundation Protocol

📌 Remember: CLOT - Clean site, Label properly, Optimal volume, Transport quickly for successful specimen processing
Specimen collection requires understanding pathogen-specific requirements, anatomical site considerations, and timing relationships with clinical presentation. Each specimen type demands specific collection techniques, transport media, and processing timelines to maintain organism viability and diagnostic accuracy.
- Blood Culture Optimization
- Collection volume: 20-30 mL total (10-15 mL per bottle set)
- Timing strategy: 2-3 sets collected 30-60 minutes apart
- Pre-antibiotic collection increases yield by 300-400%
- Optimal collection during fever spikes: 15-30 minutes before temperature peak
- Respiratory Specimens
- Sputum quality indicators: <10 epithelial cells, >25 neutrophils per low-power field
- Bronchoalveolar lavage: 100-300 mL total volume, 20 mL aliquots
- Diagnostic threshold: ≥10⁴ CFU/mL for pneumonia diagnosis
- Processing time limit: <4 hours for optimal organism recovery
- Urine Collection Standards
- Clean-catch midstream: 10-20 mL minimum volume
- Catheterized specimens: 5-10 mL adequate for analysis
- Significant bacteriuria: ≥10⁵ CFU/mL clean-catch, ≥10² CFU/mL catheterized
- Transport time: <2 hours room temperature, <24 hours refrigerated
| Specimen Type | Collection Volume | Transport Medium | Temperature | Time Limit | Critical Success Factor |
|---|---|---|---|---|---|
| Blood Culture | 20-30 mL total | Culture bottles | Room temp | <4 hours | Sterile technique |
| CSF | 2-5 mL | Sterile container | Room temp | <1 hour | Immediate processing |
| Wound/Abscess | 1-2 mL pus | Anaerobic transport | Room temp | <2 hours | Anaerobic preservation |
| Stool | Walnut-sized | Enteric transport | Room temp | <2 hours | Mucus/blood inclusion |
| Genital | Swab saturation | Specific transport | Room temp | <2 hours | Anatomical site accuracy |
💡 Master This: Understanding pre-analytical variables (collection timing, transport conditions, specimen adequacy) prevents 60-80% of diagnostic errors and false-negative results
Transport conditions critically affect organism viability and diagnostic accuracy. Different pathogens require specific environmental conditions, with fastidious organisms showing rapid viability loss under suboptimal transport conditions.
The specimen collection phase establishes the foundation for all subsequent diagnostic steps, where technical precision directly correlates with clinical diagnostic accuracy and patient care optimization.
🎯 Specimen Collection Mastery: The Foundation Protocol
⚙️ Culture Methods: The Growth Engine Architecture
📌 Remember: MAGIC - Media selection, Atmosphere control, Growth monitoring, Incubation timing, Colony characterization for successful culture
Media selection requires understanding nutritional requirements, selective properties, and differential characteristics for target organisms. Each pathogen group demands specific growth conditions, with fastidious organisms requiring enriched media and controlled atmospheric conditions.
- Primary Culture Media Categories
- Enriched media: Blood agar, Chocolate agar for fastidious organisms
- Selective media: MacConkey, Mannitol salt for specific organism groups
- Inhibitory concentrations: Crystal violet 0.0001%, Bile salts 1.5%
- Differential indicators: pH changes, hemolysis patterns, color reactions
- Atmospheric Requirements
- Aerobic incubation: 35-37°C, ambient air with 5-10% CO₂
- Anaerobic conditions: <0.5% O₂, 5-10% CO₂, 80-90% N₂
- Anaerobic indicator systems: Methylene blue, Resazurin color changes
- Incubation duration: 48-72 hours minimum for slow-growing anaerobes
- Incubation Monitoring
- Primary examination: 18-24 hours for most aerobic bacteria
- Extended incubation: 48-72 hours for fastidious organisms
- Growth detection sensitivity: 10²-10³ CFU/mL depending on media type
- Colony morphology assessment: Size, color, texture, hemolysis
| Media Type | Primary Use | Key Components | Incubation | Expected Growth | Diagnostic Yield |
|---|---|---|---|---|---|
| Blood Agar | General purpose | 5% sheep blood | 35°C, 18-24h | Most bacteria | 90-95% |
| Chocolate Agar | Fastidious organisms | Heated blood | 35°C, CO₂ | Haemophilus, Neisseria | 85-90% |
| MacConkey | Gram-negative selection | Bile salts, lactose | 35°C, 18-24h | Enterobacteriaceae | 95-98% |
| Mannitol Salt | Staphylococcus selection | 7.5% NaCl | 35°C, 24-48h | Staphylococci | 90-95% |
| Anaerobic Blood | Anaerobic bacteria | Reduced media | 35°C, 48-72h | Anaerobes | 80-85% |
💡 Master This: Understanding growth kinetics (lag phase 2-4 hours, log phase 4-12 hours, stationary phase >12 hours) enables optimal colony selection timing for accurate identification
Quality control measures ensure culture system reliability and diagnostic accuracy. Daily monitoring of media performance, incubator conditions, and growth controls maintains system integrity and prevents false-negative results.
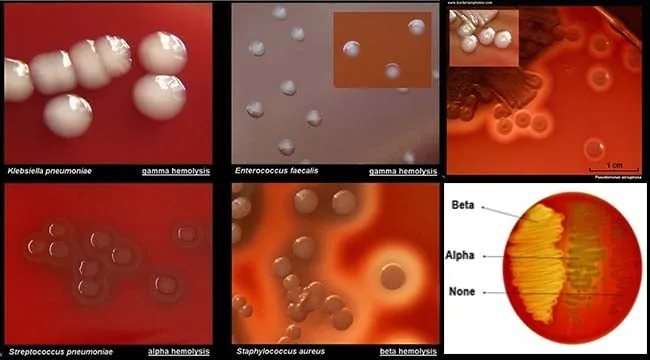
Culture methodology provides the foundation for antimicrobial susceptibility testing and serves as the reference standard against which all rapid diagnostic methods are validated.
⚙️ Culture Methods: The Growth Engine Architecture
🔍 Rapid Diagnostics: The Speed Detection Matrix
📌 Remember: RAPID - Reliable results, Accurate detection, Point-of-care capability, Immediate action, Decision support for clinical excellence
Rapid diagnostic technologies encompass immunochromatographic assays, molecular amplification methods, and automated identification systems. Each technology offers specific advantages for different clinical scenarios, with performance characteristics varying by pathogen type and specimen source.
- Immunochromatographic Assays
- Antigen detection: Streptococcus pyogenes, Influenza A/B, SARS-CoV-2
- Performance metrics: Sensitivity 80-95%, Specificity 95-99%
- Result time: 5-15 minutes for most assays
- Detection limits: 10⁵-10⁶ organisms/mL depending on target
- Molecular Rapid Tests
- PCR-based platforms: GeneXpert, FilmArray, Verigene systems
- Multiplex capabilities: 15-25 targets per panel simultaneously
- Turnaround time: 1-6 hours from specimen to result
- Analytical sensitivity: 10¹-10³ copies/mL for most targets
- Automated Identification
- MALDI-TOF mass spectrometry: Organism ID in 2-5 minutes
- Automated susceptibility: 4-18 hours for complete antibiogram
- Database coverage: >2,000 organisms with >95% accuracy
- Cost per test: $2-5 compared to $15-25 conventional methods
| Technology | Time to Result | Sensitivity | Specificity | Targets per Test | Clinical Application |
|---|---|---|---|---|---|
| Antigen Tests | 5-15 minutes | 80-95% | 95-99% | 1-2 | Point-of-care screening |
| Molecular Panels | 1-6 hours | 95-99% | 98-100% | 15-25 | Syndromic diagnosis |
| MALDI-TOF | 2-5 minutes | 85-95% | >95% | ID only | Rapid identification |
| Automated AST | 4-18 hours | 90-98% | 95-99% | 15-20 drugs | Resistance detection |
| Biosensors | 30-60 minutes | 85-95% | 90-98% | 1-5 | Critical care monitoring |
💡 Master This: Understanding analytical sensitivity versus clinical sensitivity prevents over-reliance on rapid tests with detection limits that may miss low-level infections requiring culture confirmation
Cost-effectiveness analysis demonstrates rapid testing value through reduced length of stay, optimized antibiotic use, and improved patient outcomes. Integration with antimicrobial stewardship programs maximizes clinical impact while controlling healthcare costs.
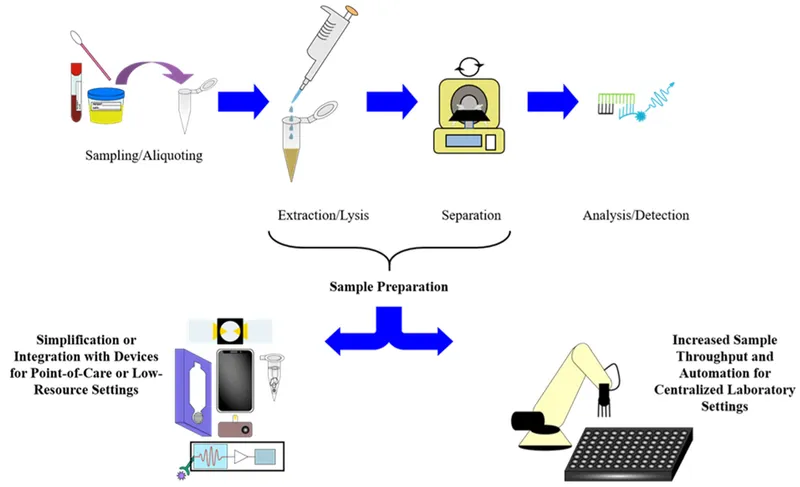
Rapid diagnostics complement rather than replace traditional culture methods, creating comprehensive diagnostic algorithms that optimize both speed and accuracy for superior patient care.
🔍 Rapid Diagnostics: The Speed Detection Matrix
⚖️ Antimicrobial Susceptibility: The Resistance Intelligence Network

📌 Remember: CLEARS - Clinical breakpoints, Laboratory standards, Error detection, Accuracy controls, Resistance mechanisms, Stewardship support
Susceptibility testing employs multiple methodologies, each with specific applications, performance characteristics, and clinical utility. Understanding method selection criteria and result interpretation enables optimal integration of susceptibility data with clinical decision-making processes.
- Disk Diffusion Method (Kirby-Bauer)
- Zone diameter measurements: Susceptible, Intermediate, Resistant categories
- Quality control ranges: ±2-3 mm for standard control strains
- E. coli ATCC 25922: Ampicillin 16-22 mm, Ciprofloxacin 30-40 mm
- S. aureus ATCC 25923: Oxacillin 18-24 mm, Vancomycin 17-21 mm
- Broth Microdilution
- MIC determination: Minimum Inhibitory Concentration in μg/mL
- Automated systems: 96-384 well formats with 2-fold dilutions
- Reading time: 16-20 hours for most organisms
- Accuracy: >95% agreement with reference methods
- Gradient Methods (E-test)
- MIC strips: Continuous concentration gradients on agar surface
- Applications: Fastidious organisms, Special antibiotics
- Cost factor: 3-5x higher than disk diffusion
- Correlation: >90% agreement with broth microdilution
| Method | Result Type | Time Required | Cost per Test | Automation Level | Clinical Application |
|---|---|---|---|---|---|
| Disk Diffusion | Zone diameter | 18-24 hours | $2-3 | Manual reading | Routine screening |
| Broth Microdilution | MIC value | 16-20 hours | $5-8 | Automated | Precise dosing |
| Gradient Strip | MIC value | 18-24 hours | $8-12 | Manual reading | Fastidious organisms |
| Rapid Systems | MIC/Category | 4-8 hours | $10-15 | Fully automated | Critical infections |
| Molecular | Gene detection | 2-6 hours | $15-25 | Automated | Resistance prediction |
💡 Master This: Understanding pharmacokinetic/pharmacodynamic relationships enables interpretation of MIC values relative to achievable tissue concentrations for optimal therapeutic outcomes
Resistance mechanism detection enhances susceptibility testing by identifying specific resistance genes and predicting phenotypic resistance patterns. Molecular methods provide rapid resistance detection, particularly for slow-growing organisms and critical resistance markers.

Quality assurance programs ensure susceptibility testing accuracy through daily control strain testing, proficiency testing participation, and systematic error detection protocols that maintain clinical reliability.
⚖️ Antimicrobial Susceptibility: The Resistance Intelligence Network
🔗 Molecular Integration: The Genetic Code Decoder
📌 Remember: PRIMER - Pathogen targets, Resistance genes, Identification speed, Multiplex capability, Error prevention, Result validation
Molecular methods encompass various amplification technologies, each offering specific advantages for different clinical applications. Understanding technology selection criteria and performance characteristics enables optimal test utilization for diverse diagnostic scenarios.
- PCR-Based Amplification
- Real-time PCR: Quantitative results with Ct values indicating organism load
- Multiplex PCR: 15-25 targets simultaneously in single reaction
- Analytical sensitivity: 1-100 copies/mL for most viral targets
- Specificity: >99% with proper primer design and validation
- Isothermal Amplification
- LAMP technology: Constant temperature amplification at 60-65°C
- Point-of-care applications: 30-60 minutes without thermal cycling
- Equipment requirements: Minimal compared to PCR systems
- Cost per test: $5-10 versus $15-25 for PCR methods
- Next-Generation Sequencing
- Metagenomic analysis: Unbiased pathogen detection from clinical specimens
- Resistance profiling: Complete resistome characterization
- Turnaround time: 24-48 hours for comprehensive analysis
- Data output: Gigabytes requiring bioinformatics expertise
| Technology | Targets per Test | Time to Result | Sensitivity | Quantitative | Clinical Application |
|---|---|---|---|---|---|
| Real-time PCR | 1-4 | 2-4 hours | 95-99% | Yes | Targeted detection |
| Multiplex PCR | 15-25 | 1-6 hours | 90-98% | Limited | Syndromic panels |
| LAMP | 1-3 | 30-60 minutes | 85-95% | No | Point-of-care |
| Sequencing | Unlimited | 24-48 hours | >95% | Yes | Unknown pathogens |
| Microarray | 50-100 | 4-8 hours | 90-95% | No | Resistance profiling |
💡 Master This: Understanding analytical sensitivity (copies/mL) versus clinical sensitivity (correlation with disease) prevents over-interpretation of positive molecular results in asymptomatic patients
Molecular testing limitations include inability to assess organism viability, potential for contamination, and requirement for antimicrobial susceptibility testing by alternative methods. Integration with culture methods provides comprehensive diagnostic approaches combining speed with complete therapeutic guidance.
Quality control measures for molecular testing include extraction controls, amplification controls, and inhibition monitoring to ensure accurate results and prevent false-negative reporting due to technical failures.
🔗 Molecular Integration: The Genetic Code Decoder
🎯 Clinical Integration Arsenal: The Diagnostic Command Center
📌 Remember: IMPACT - Integrated results, Multiple methods, Patient correlation, Antibiotic guidance, Clinical outcomes, Timing optimization
Clinical integration requires understanding the complementary roles of different diagnostic methods, their relative strengths and limitations, and optimal sequencing for specific clinical scenarios. Effective integration maximizes diagnostic accuracy while optimizing resource utilization and patient care efficiency.
- Diagnostic Algorithm Integration
- Rapid screening: Rule-out negative results with >95% NPV
- Culture confirmation: Definitive identification and susceptibility testing
- Turnaround optimization: Rapid results in 2-6 hours, Complete data in 24-48 hours
- Cost-effectiveness: $50-100 comprehensive workup versus $200-500 empirical therapy costs
- Clinical Correlation Framework
- Pre-test probability: Clinical presentation scoring systems
- Post-test interpretation: Likelihood ratios for positive and negative results
- Sensitivity integration: Sequential testing improves overall sensitivity to >98%
- Specificity maintenance: Confirmatory testing prevents false-positive treatment decisions
- Antimicrobial Stewardship Integration
- Rapid de-escalation: Narrow-spectrum therapy based on definitive results
- Duration optimization: Biomarker-guided therapy discontinuation
- Antibiotic reduction: 20-40% decrease in broad-spectrum usage
- Resistance prevention: Targeted therapy reduces selection pressure
| Clinical Scenario | Primary Method | Confirmatory Test | Time to Action | Diagnostic Accuracy | Cost Optimization |
|---|---|---|---|---|---|
| Sepsis Workup | Blood culture + Rapid PCR | Susceptibility testing | 2-6 hours | >95% | 30-40% reduction |
| Pneumonia Diagnosis | Respiratory culture + Antigen | Molecular panel | 4-24 hours | 90-95% | 25-35% reduction |
| UTI Evaluation | Urine culture + Rapid screen | Automated susceptibility | 1-18 hours | >98% | 40-50% reduction |
| Wound Infection | Culture + Gram stain | Anaerobic culture | 18-72 hours | 85-95% | 20-30% reduction |
| Meningitis Workup | CSF culture + Molecular | Antigen detection | 1-24 hours | >95% | High priority |
💡 Master This: Understanding diagnostic stewardship principles enables optimal test selection that maximizes clinical value while minimizing unnecessary testing and healthcare costs
Quality metrics for diagnostic integration include turnaround time monitoring, clinical correlation assessment, and outcome tracking to ensure optimal patient care delivery. Continuous improvement processes optimize diagnostic workflows and enhance clinical decision support capabilities.

Effective diagnostic integration requires multidisciplinary collaboration between laboratory professionals, clinicians, and antimicrobial stewardship teams to optimize patient outcomes through evidence-based diagnostic and therapeutic decision-making.
🎯 Clinical Integration Arsenal: The Diagnostic Command Center
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app
























