Amino Acids - The Alpha Team
- Building blocks of proteins. General structure: central $\alpha$-carbon, amino group ($-NH_2$), carboxyl group ($-COOH$), H atom, & variable R-group (side chain).
- All (except Glycine) are chiral; L-isomers predominate in humans.
- Classified by R-group properties: Nonpolar, Aromatic, Polar uncharged, Positively charged (Basic), Negatively charged (Acidic).
- Essential AAs (dietary intake crucial): 📌 PVT TIM HALL (Phenylalanine, Valine, Threonine, Tryptophan, Isoleucine, Methionine, Histidine, Arginine, Leucine, Lysine).
- At physiological pH (approx. 7.4), exist as zwitterions (dipolar ions).
- Peptide bond: Covalent bond linking amino acids; formed by dehydration synthesis (loss of $H_2O$).

⭐ Aromatic amino acids (Tyrosine, Tryptophan, Phenylalanine) are responsible for UV light absorption by proteins, primarily at 280 nm (Tyr & Trp).
Protein Structure - Levels Unlocked
- Primary (1°): Linear sequence of amino acids.
- Linked by peptide bonds.
- Determines all higher structures; read N-terminus → C-terminus.
- Example: Insulin.
- Secondary (2°): Local, regular folding of polypeptide backbone.
- Stabilized by hydrogen bonds between backbone CO & NH groups.
- α-helix: Right-handed coil; 3.6 residues/turn. (📌 Proline = Helix Breaker).
- β-pleated sheet: Extended strands; parallel or antiparallel.
- β-turns & loops: Connect α-helices and β-sheets.
- Tertiary (3°): Overall 3D conformation of a single polypeptide chain.
- Maintained by R-group (side chain) interactions:
- Hydrophobic interactions (major driving force).
- Hydrogen bonds.
- Ionic bonds (salt bridges).
- Disulfide bonds (covalent; Cysteine-Cysteine).
- Forms functional domains.
- Maintained by R-group (side chain) interactions:
- Quaternary (4°): Arrangement of multiple polypeptide subunits (oligomeric proteins).
- Held by non-covalent interactions & sometimes disulfide bonds.
- Example: Hemoglobin (tetramer: 2α, 2β subunits).
⭐ The primary structure, the specific amino acid sequence, is the ultimate determinant of a protein's three-dimensional conformation and its biological function.

Protein Folding - Shape Shifters
- Process: Linear polypeptide chain → functional 3D structure. Primary sequence dictates folding.
- Chaperones (e.g., HSPs): Aid correct folding, prevent misfolding & aggregation.
- HSP60 (chaperonins): Folding chambers.
- HSP70: Bind nascent polypeptides.
- Misfolding consequences:
- Degradation by Ubiquitin-Proteasome System (UPS).
- Aggregation → Amyloid fibrils → Disease.
- Key Diseases: Alzheimer's (Aβ), Parkinson's (α-synuclein), Prion diseases (PrPSc), Cystic Fibrosis (CFTR).

⭐ Prion protein (PrP) misfolding into PrPSc (scrapie form) causes transmissible spongiform encephalopathies (e.g., CJD); PrPSc acts as a template for converting normal PrPC.
Hemoglobin & Myoglobin - Oxygen's Ride
- Hemoglobin (Hb):
- Structure: Tetramer (α₂β₂ in HbA), 4 hemes, binds 4 O₂.
- Function: O₂ transport in blood. Sigmoidal O₂ dissociation curve (ODC) due to cooperative binding.
- Regulation (Right shift ODC = ↑O₂ unloading): ↑ 2,3-BPG, ↑ $PCO_2$ (Bohr), ↑ H⁺ (Bohr, ↓pH), ↑ Temp. 📌 CADET right shift!
- Myoglobin (Mb):
- Structure: Monomer, 1 heme, binds 1 O₂.
- Function: O₂ storage in muscle. Hyperbolic ODC, higher O₂ affinity than Hb.
- Fetal Hb (HbF - α₂γ₂): Higher O₂ affinity (poor 2,3-BPG binding) facilitates placental O₂ transfer.
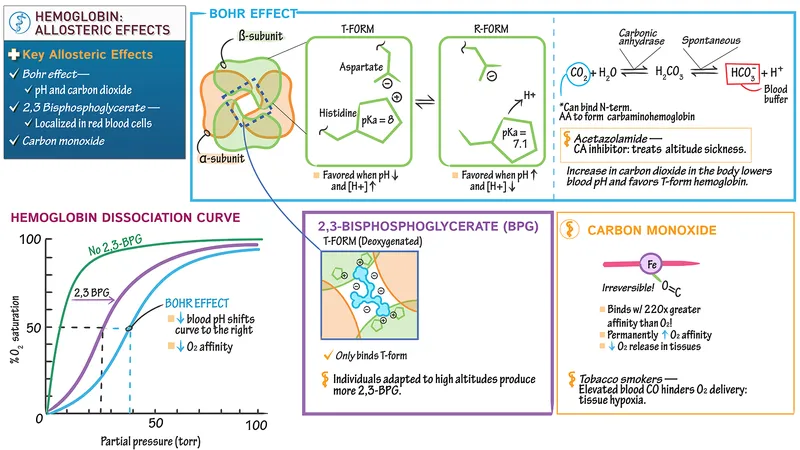
⭐ Carbon monoxide (CO) binds Hb with ~200-250 times greater affinity than O₂, forming carboxyhemoglobin (COHb), impairing O₂ transport and causing a left shift of the remaining Hb's ODC.
High‑Yield Points - ⚡ Biggest Takeaways
- Primary structure (amino acid sequence) dictates higher-order structures and function.
- Secondary structures (α-helix, β-sheet) are stabilized by hydrogen bonds.
- Tertiary structure (3D folding) is stabilized by hydrophobic interactions, disulfide bonds, and ionic bonds.
- Quaternary structure involves multiple polypeptide subunits (e.g., hemoglobin).
- Protein misfolding can cause diseases like Alzheimer's (amyloid-β) and prion diseases.
- Enzymes (protein catalysts) lower activation energy, speeding reactions.
- Allosteric regulation involves binding at a site distinct from the active site, altering protein activity.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

