Principle & Basics - Microbial Fingerprinter
- MALDI-TOF MS (Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry): Rapid, accurate microbial identification based on unique protein (mainly ribosomal) profiles.
- Core Principle: "Soft" ionization of whole microbial cells or extracts, generating a characteristic mass spectrum (fingerprint).
- Matrix: Sample mixed with matrix (e.g., α-cyano-4-hydroxycinnamic acid); co-crystallizes.
- Laser Desorption/Ionization: Pulsed laser desorbs and ionizes matrix & sample molecules (mostly singly charged, [M+H]$^+$).
- Time-of-Flight (TOF) Analyzer: Ions accelerated into a flight tube. Smaller ions travel faster.
- $t = d \sqrt{m/(2zV)}$, where $t$=time, $d$=distance, $m$=mass, $z$=charge, $V$=voltage.
- Detector: Records ion arrival times.
- Mass Spectrum: Plot of ion intensity vs. mass-to-charge ratio (m/z), creating a unique "fingerprint".
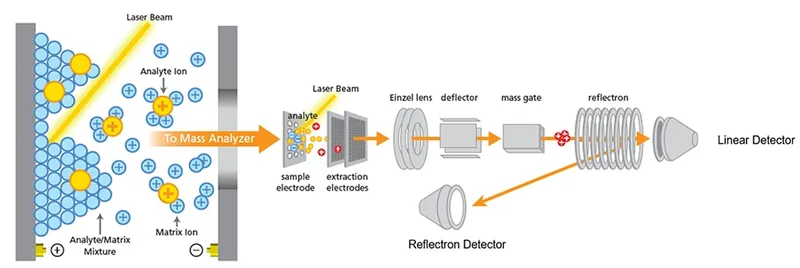
⭐ MALDI-TOF MS primarily analyzes ribosomal proteins due to their abundance and conservation within species, yet sufficient variability between species for discrimination. This makes them ideal biomarkers for microbial identification.
Workflow & Steps - Sample to Spectrum
Rapid microbial ID via protein fingerprinting (mainly ribosomal proteins).
- 1. Sample Prep:
- Direct Colony Transfer (DCT): Pure colony smeared on MALDI plate.
- Extraction: For yeasts, mycobacteria; uses formic acid/acetonitrile for protein release.
- 2. Matrix & Co-crystallization:
- Sample + Matrix (e.g., HCCA - α-cyano-4-hydroxycinnamic acid).
- Air-dry: Forms sample-matrix co-crystals.
- 3. MALDI-TOF MS:
- Laser Desorption/Ionization: UV laser hits crystals → gas-phase ions (mostly proteins).
- Ion Acceleration: Electric field accelerates ions into flight tube.
- Time-of-Flight (TOF): Ions separated by mass-to-charge ($m/z$); smaller ions faster.
- 4. Spectrum & ID:
- Mass spectrum generated (peak pattern = fingerprint).
- Compared to database for species identification.
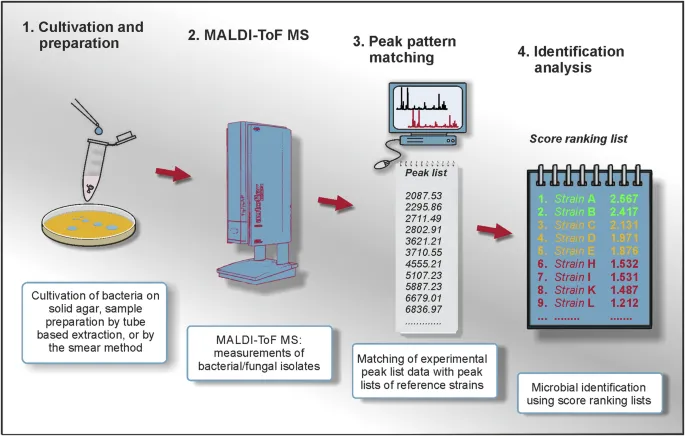
⭐ The most abundant ions detected in MALDI-TOF MS of bacteria are ribosomal proteins, which are conserved yet species-specific, providing a reliable "fingerprint".
Clinical Applications - Microbe ID Ace
- Core Function: Rapid, precise identification of cultured microorganisms.
- Bacteria: Wide range (aerobes, anaerobes, Gram +ve/-ve).
- Yeasts: Common species (e.g., Candida spp.).
- Filamentous Fungi: Expanding databases.
- Mycobacteria: With specific extraction protocols.
- Specimen Versatility:
- Colonies from culture media (solid/liquid).
- Directly from positive blood culture broths (crucial for sepsis).
- Impact on Workflow:
- Speed: Identification in minutes.
- Accuracy: High for routine isolates.
- Efficiency: Reduces reliance on numerous biochemical tests.
- Developing Frontiers:
- Rapid Antimicrobial Susceptibility Testing (AST) (e.g., β-lactamase detection).
- Epidemiological strain typing.
⭐ MALDI-TOF MS enables identification of bacteria and yeasts directly from positive blood cultures, often within 15-30 minutes of a positive signal.
Advantages & Limitations - Balancing Act
- Advantages:
- Speed: Rapid ID (minutes vs. hours/days).
- Accuracy: High for common, well-characterized isolates.
- Cost-Effective: Low consumable cost; ↓ long-term lab costs.
- Automation: Suitable for high-throughput workflows.
- Minimal Sample Prep: Direct colony testing often feasible.
- Limitations:
- Database Dependency: Performance relies on database quality & updates.
- Initial Cost: Significant capital investment.
- Differentiation: Struggles with closely related species/subspecies.
- Culture Requirement: Needs isolated colonies, not direct polymicrobial samples.
- No Susceptibility Data: Cannot provide antimicrobial resistance (AST).
⭐ MALDI-TOF identifies microbes via their unique protein fingerprint, mainly ribosomal proteins.
High‑Yield Points - ⚡ Biggest Takeaways
- MALDI-TOF MS identifies microbes by analyzing ribosomal protein profiles.
- It measures the mass-to-charge ratio (m/z) of these proteins.
- A matrix (e.g., HCCA) is crucial for desorption and ionization.
- Time-of-Flight (TOF) analyzer separates ions based on their travel time.
- Enables rapid identification of bacteria and fungi, often to species level.
- Database comparison of protein spectra is key for identification.
- Limited for direct specimen analysis; typically requires prior culture.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

