Enzyme Kinetics: Basics - Speedy Catalysts
- Catalysts (proteins/RNA) that ↑ reaction rates.
- Mechanism: Lower activation energy ($E_a$).
- Provide alternative reaction pathway.
- Stabilize transition state.
- Crucially, enzymes do NOT alter $\Delta G$ (Gibbs free energy) or $K_{eq}$ (equilibrium constant).
- Reaction Velocity (V): Rate of [product] formation or [substrate] consumption.
- Factors Affecting Velocity:
- Enzyme concentration [E]: V ∝ [E] (if [S] saturating).
- Substrate concentration [S]: V ↑ with [S] until $V_{max}$.
- Temperature: Bell-shaped curve (optimum T); high T denatures.
- pH: Bell-shaped curve (optimum pH); extreme pH denatures.
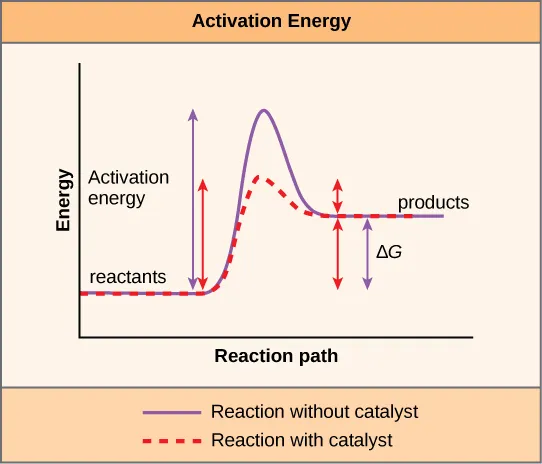
⭐ While enzymes don't change the reaction's endpoint (equilibrium), they drastically reduce the time taken to reach it.
Enzyme Kinetics: Michaelis-Menten - The Speed Formula
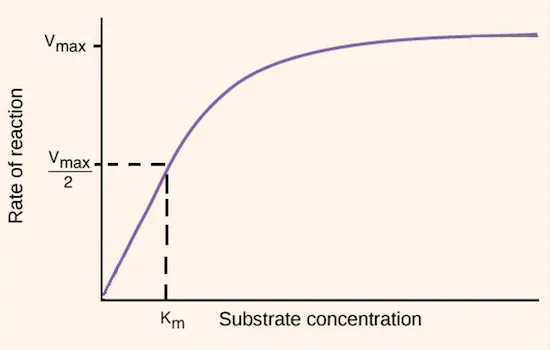
- Describes how initial reaction velocity (V) changes with substrate concentration ([S]).
- The fundamental equation: $V = \frac{V_{max}[S]}{K_m + [S]}$
- $V_{max}$ (Maximum Velocity):
- The theoretical maximum rate when all enzyme active sites are saturated.
- Directly proportional to total enzyme concentration [E]$_{total}$; units: rate (e.g., µmol/min).
- $K_m$ (Michaelis Constant):
- Substrate concentration ([S]) at which V = ½ $V_{max}$.
- An inverse measure of enzyme's affinity for its substrate: ↓ $K_m$ implies ↑ affinity.
- Units: concentration (e.g., M, mM).
- Key Assumptions:
- [S] >> [E] (substrate in large excess).
- Steady-state: d[ES]/dt = 0 (ES complex concentration is constant).
- Initial velocities measured (product P ≈ 0, reverse reaction is negligible).
⭐ Low $K_m$ indicates the enzyme is highly efficient at low substrate concentrations.
Enzyme Kinetics: Lineweaver-Burk - Straight Plot Story
- A double reciprocal plot of Michaelis-Menten kinetics: linearizes data.
- Equation: $\frac{1}{V_0} = \left(\frac{K_m}{V_{max}}\right) \frac{1}{[S]} + \frac{1}{V_{max}}$ (form $y=mx+c$)
- Plot parameters:
- Y-axis: $\frac{1}{V_0}$
- X-axis: $\frac{1}{[S]}$
- Y-intercept: $\frac{1}{V_{max}}$
- X-intercept: $-\frac{1}{K_m}$
- Slope: $\frac{K_m}{V_{max}}$
- 📌 Mnemonic for intercepts: Y-intercept is $\frac{1}{V_{max}}$ (V for Vertical). X-intercept is $-\frac{1}{K_m}$ (negative value on X-axis).
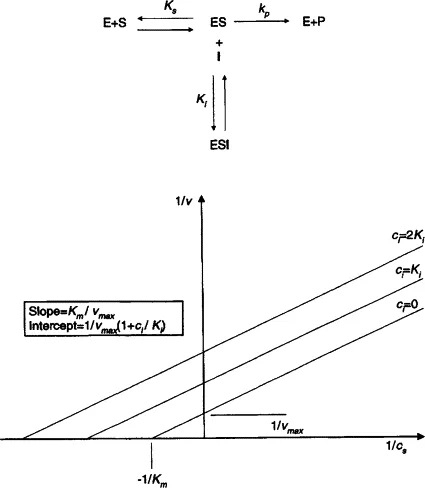
- Key for determining $K_m$, $V_{max}$, and analyzing enzyme inhibition types.
- Limitation: Distorts errors at low [S] (high $\frac{1}{[S]}$ values), less accurate.
⭐ Lineweaver-Burk plots clearly distinguish competitive, non-competitive, and uncompetitive inhibition patterns by their effects on $V_{max}$ and $K_m$.
Enzyme Kinetics: Inhibition - Kinetic Blockers
- Competitive Inhibition:
- Inhibitor (I) mimics substrate; binds active site.
- Effect: $K_m$ ↑ (affinity ↓), $V_{max}$ unchanged.
- Reversible by ↑ [S].
- E.g., Statins, Methotrexate.
- Non-competitive Inhibition:
- I binds E or ES at an allosteric site.
- Effect: $K_m$ unchanged, $V_{max}$ ↓.
- Not reversible by ↑ [S].
- E.g., Lead, some forms of Cyanide.
- Uncompetitive Inhibition:
- I binds only to ES complex (allosteric site).
- Effect: Both $K_m$ ↓ and $V_{max}$ ↓ (proportionally).
- Lineweaver-Burk: Parallel lines.
- E.g., Lithium.
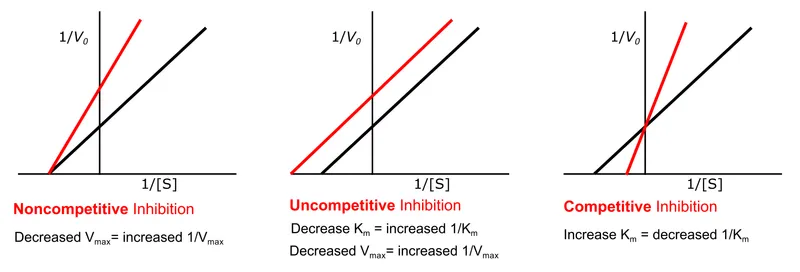
⭐ Competitive inhibitors increase the Michaelis constant (apparent $K_m$) but do not affect the maximum velocity ($V_{max}$). More substrate is needed to reach half $V_{max}$, but $V_{max}$ itself is still achievable with sufficient substrate concentration.
High‑Yield Points - ⚡ Biggest Takeaways
- Michaelis-Menten equation: Relates initial velocity (v₀) to substrate concentration [S].
- K_m_ (Michaelis constant): [S] at ½ V_max_. Inverse measure of substrate affinity (↓K_m_ = ↑affinity).
- V_max_ (maximum velocity): Maximum velocity at enzyme saturation. Proportional to enzyme concentration.
- Lineweaver-Burk plot: Double reciprocal (1/v vs 1/[S]), linearizes kinetics.
- Competitive inhibitors: ↑K_m_, V_max_ unchanged. Overcome by ↑[S].
- Non-competitive inhibitors: ↓V_max_, K_m_ unchanged. Not overcome by ↑[S].
- Uncompetitive inhibitors: ↓V_max_ and ↓K_m_. Bind ES complex only.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

