PopGen Forensics: Basics & HWE - Gene Pool Rules
- Population Genetics: Studies genetic variation in populations.
- Gene Pool: Total genetic information in an interbreeding population.
- Allele Frequency ($p, q$): Proportion of a specific allele. Formula: $p + q = 1$.
- Genotype Frequency ($p^2, 2pq, q^2$): Proportion of a specific genotype.
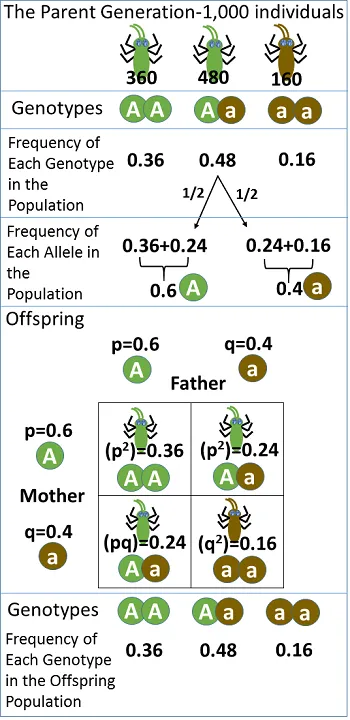
- Hardy-Weinberg Equilibrium (HWE): States allele and genotype frequencies remain stable if specific conditions are met.
- Equation: $p^2 + 2pq + q^2 = 1$.
- Assumptions (📌 LRM-NoMMS):
- Large population
- Random Mating
- No Mutation
- No Migration (gene flow)
- No Selection
- Forensic Significance: Essential for estimating DNA profile rarity and calculating statistical match probability.
⭐ HWE is key to calculating the Random Match Probability (RMP), indicating the likelihood of an unrelated individual matching a DNA profile by chance.
PopGen Forensics: Allele Frequencies & Databases - Counting Our Genes
- Allele Frequency Estimation: Crucial for statistical weight. Calculated by counting specific alleles within a sample population. E.g., $p = (\text{count of allele A}) / (2 \times \text{total individuals})$.
- Population Databases: Store DNA profiles (typically STR loci) to estimate these frequencies.
- International examples: CODIS (Combined DNA Index System - USA), ENFSI (European Network of Forensic Science Institutes).
- National: Indian-specific databases vital for local relevance, though comprehensive representative databases remain challenging due to population diversity.
- Key Loci: Primarily Short Tandem Repeats (STRs); Variable Number Tandem Repeats (VNTRs) have niche applications in specific contexts.
- Database Integrity:
- Must be representative of the target population.
- Challenges: Ethnic/population stratification, adequate database size, accounting for sub-population differences.
⭐ CODIS mandates analysis of 20 core STR loci (expanded from 13 in 2017) for consistent DNA profile comparison and searches across forensic databases in the USA.
PopGen Forensics: Statistical Interpretation - Odds On Identity
- RMP (Random Match Probability): Chance of coincidental profile match. Modern probabilistic genotyping software (e.g., STRmix) directly calculates LRs without simplified RMP approximations.
- Traditional formulas: Homozygotes $p^2$; Heterozygotes $2pq$ (now largely superseded).
- LR (Likelihood Ratio): $LR = P(E|H_p) / P(E|H_d)$. Compares prosecution ($H_p$) vs. defense ($H_d$) hypotheses. Probabilistic systems provide robust LR calculations. Higher LR = stronger evidence for $H_p$.
- Population Substructure: Advanced statistical tools account for population complexities more accurately than manual corrections:
- Traditional Theta (θ) Correction: Homozygote $p^2 + p(1-p)\theta$; Heterozygote $2pq(1-\theta)$.
- Modern software integrates these corrections automatically.
- ⚠️ Statistical Interpretation Focus:
- Proper LR Application: Understanding limitations of statistical interpretations under BSA (Bharatiya Sakshya Adhiniyam 2023) evidence standards.
- Robust Frameworks: Probabilistic genotyping mitigates interpretation complexities in BNSS proceedings.
- Bayesian Statistics: Integrates LR with prior odds: Prior Odds $\times$ LR = Posterior Odds.
⭐ LR is key in DNA interpretation, comparing $P(E|H_p)$ vs $P(E|H_d)$. An LR of 1000 means evidence is 1000x more likely if suspect is source vs. random person.
PopGen Forensics: Genetic Variation Factors - Shifting Gene Sands
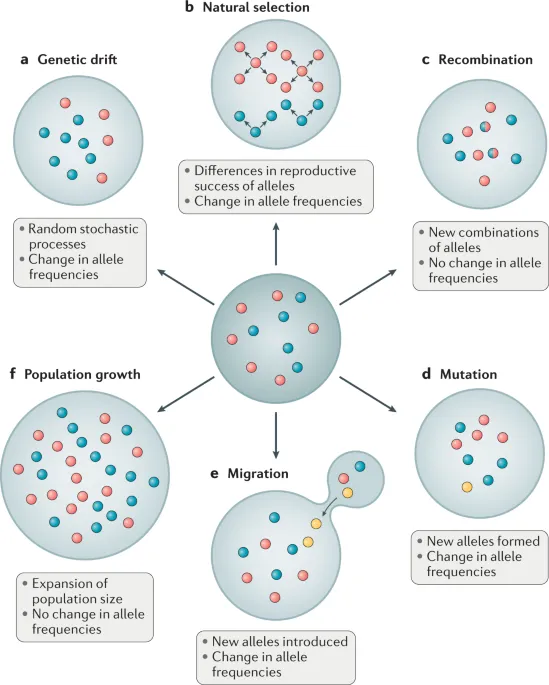
Factors altering allele frequencies from Hardy-Weinberg Equilibrium (HWE) expectations:
- Mutation: Ultimate source of new alleles; generally low rate for STRs.
- Gene Flow (Migration): Movement of alleles between populations; homogenizes allele frequencies, impacts database representativeness.
- Genetic Drift: Random fluctuations in allele frequencies, more pronounced in small populations.
- Bottleneck Effect: Drastic reduction in population size (e.g., disaster).
- Founder Effect: New population established by a small number of individuals.
- Natural Selection: Differential survival/reproduction (minimal impact on most forensic STR markers).
- Population Substructure: Existence of distinct subgroups within a larger population.
- Wahlund Effect: Reduction in observed heterozygosity and excess homozygosity if subpopulations with different allele frequencies are pooled and treated as a single random-mating unit.
- Linkage Disequilibrium (LD): Non-random association of alleles at different loci; relevant for haplotype frequency estimation.
⭐ The Wahlund effect is a key consideration in forensic genetics, as failure to account for population substructure can lead to inaccurate statistical interpretations of DNA evidence, particularly underestimating match probabilities for rare genotypes if a general database is used without considering specific sub-population frequencies.
High‑Yield Points - ⚡ Biggest Takeaways
- Hardy-Weinberg Equilibrium (HWE) provides foundational concepts, but modern forensic DNA analysis increasingly uses probabilistic genotyping software (PGS) for complex samples and NGS applications.
- Likelihood ratios (LRs) from PGS are replacing simple Random Match Probability (RMP) calculations, especially for complex DNA mixtures and low-template DNA.
- Comprehensive population databases with enhanced genetic diversity and AI integration improve statistical accuracy beyond traditional ethnicity-based classifications.
- Unlinked STR markers and advanced statistical software effectively mitigate linkage disequilibrium complications in routine forensic analysis.
- Database validation, transparency, and interoperability of national DNA databases enhance forensic reliability under BSA provisions.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

