Extraction Principles - Code Cracking Intro
- Nucleic acids (DNA/RNA): Microbial genetic material; life's code.
- Goal: Isolate pure, intact DNA or RNA from microbes.
- Essential for: Diagnostics (PCR), sequencing, research, epidemiology.
- Challenges:
- Efficient cell lysis (e.g., tough bacterial walls).
- Preventing nuclease degradation.
- Removing assay inhibitors (e.g., heme, polysaccharides).
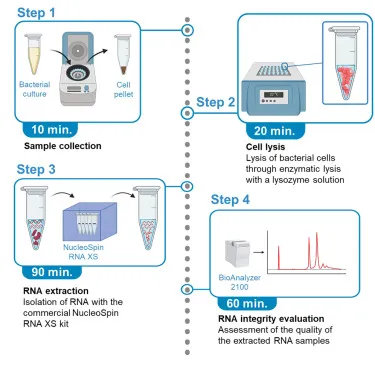
⭐ Purity (A260/A280 ratio: DNA ~1.8, RNA ~2.0) and integrity are critical for downstream applications like PCR success.
General Steps - The Great Unraveling
📌 Mnemonic: Large Purple Pandas Want Eucalyptus (Lysis, Purification, Precipitation, Wash, Elution).
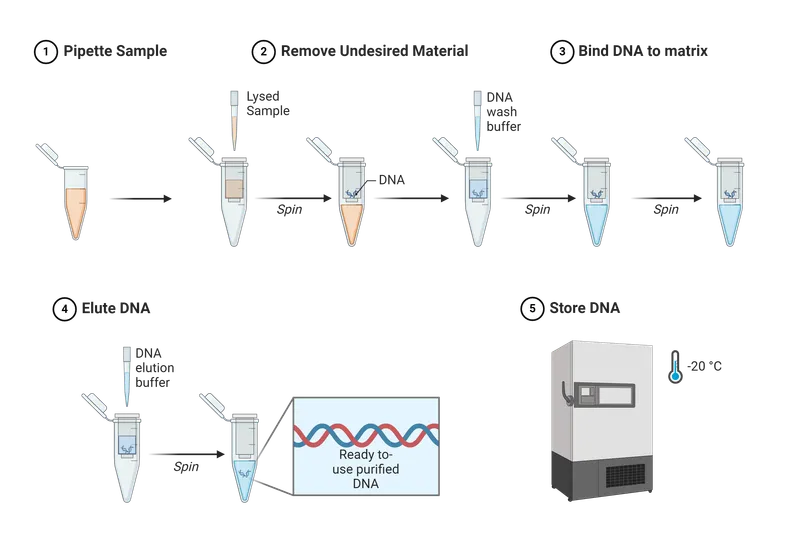
- Lysis (Cell Breakage): Key to release NA.
- Mechanical: Beads, sonication.
- Chemical: Detergents (SDS), chaotropic salts (guanidinium thiocyanate).
- Enzymatic: Lysozyme (bacteria), Lyticase (yeast), Proteinase K (degrades proteins/nucleases).
- Purification (Remove Contaminants): Isolate NA from cellular debris.
- Organic extraction: Phenol-chloroform.
- Solid-phase: Silica columns (NA binds in high salt, elutes in low salt).
- Concentration & Wash: Concentrate NA and remove residual impurities.
- Precipitation: Cold ethanol or isopropanol + salt.
- Wash: 70% ethanol (removes salts/contaminants).
- Elution (Resuspend NA): Rehydrate purified NA.
- Nuclease-free $H_2O$ or TE buffer.
⭐ Proteinase K is crucial; it degrades most proteins, including nucleases that would otherwise degrade DNA/RNA, and remains active in detergents like SDS and chaotropic salts during lysis and purification steps.
DNA Methods - Blueprint Retrievers
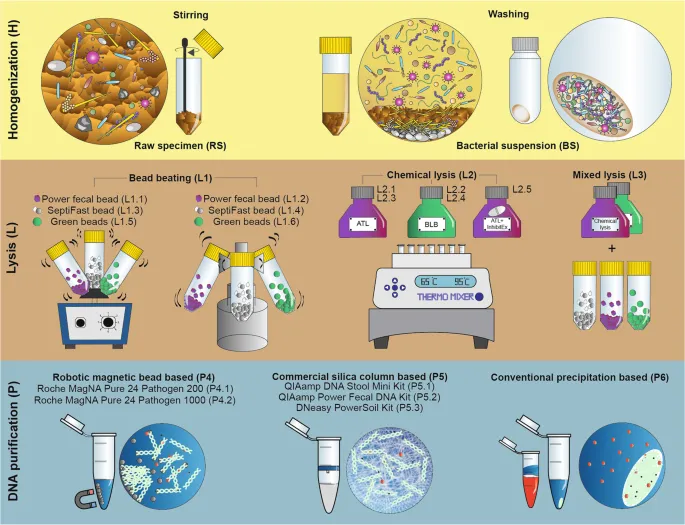
- Goal: Isolate pure DNA from microbes for PCR, sequencing.
- Core Steps:
- Cell Lysis: Break cells.
- Contaminant Removal (proteins, RNA).
- DNA Recovery.
- Lysis Types:
- Mechanical: Bead beating, sonication. Physical force. Use: Tough cells (fungi, Gram+). Pro: Effective. Con: DNA shear.
- Chemical: Detergents (SDS), chaotropes. Membrane disruption. Pro: Gentler. Con: Inhibitors.
- Enzymatic: Lysozyme (bacteria), lyticase (yeast), Proteinase K (protein digestion, nuclease inactivation). Specific. Pro: Mild. Con: Cost. 📌 Pro-K for Nuclease Knell!
- Purification:
- Phenol-Chloroform (PCI): Phase separation. DNA in aqueous. ⚠️ Toxic.
- Silica Column: DNA binds silica (high salt), elutes (low salt). Fast, pure.
⭐ Silica columns offer rapid, high-purity DNA ideal for sensitive assays like qPCR & NGS.
RNA & QC - Messenger Wrangling
- Challenge: Ubiquitous, stable RNases degrade RNA. 📌 RNA Needs Always Careful Extraction (RNACE).
- RNase Control:
- Strict RNase-free environment (gloves, consumables).
- Chemicals: DEPC (water/buffers), GITC (lysis buffer, Trizol).
- Inhibitors: Proteinaceous (e.g., RNasin®).
- RNA Extraction:
- Methods: Organic (Phenol-GITC) or Solid-phase (silica).
- DNase I treatment: Essential for RT-PCR, RNA-seq (removes gDNA).
- RNA Quality Control (QC):
- Purity (A260/A280): Target ~2.0. (<1.8 = protein contam.).
- Purity (A260/A230): Target 1.8-2.2. (<1.8 = phenol/salt contam.).
- Integrity (Gel): 28S & 18S rRNA (eukaryotes), ratio ~2:1. Smear = degradation.
- Integrity (RIN): Scale 1-10. >7 good; >8 for NGS/sensitive assays.
- Concentration: A260nm or fluorometry (Qubit for specificity).
⭐ High-quality RNA for demanding applications (NGS, microarrays) requires A260/A280 ≈ 2.0, A260/A230 >2.0, and RIN >8.

High‑Yield Points - ⚡ Biggest Takeaways
- Phenol-chloroform extraction is classic but toxic; separates by phase.
- Silica spin columns ensure rapid, pure DNA/RNA isolation using chaotropic salts.
- Alkaline lysis is standard for bacterial plasmid DNA extraction.
- Enzymatic lysis (lysozyme, proteinase K) is vital for cell disruption.
- RNA extraction demands RNase inhibitors (e.g., DEPC) or guanidinium thiocyanate.
- Automated systems offer high-throughput, standardized nucleic acid isolation.
- Method choice depends on sample, target nucleic acid (DNA/RNA), and application.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

