Microarray Basics - Chips & Dips
- Core Principle: Massively parallel hybridization. Labeled sample nucleic acids (targets) bind to complementary, immobilized oligonucleotide probes on a solid surface (chip).
- Key Components:
- Microarray Chip: Solid support (e.g., glass, silicon) with thousands of spots.
- Probes: Short, known DNA/RNA sequences (oligonucleotides, cDNA) fixed to chip spots.
- Target: Fluorescently labeled nucleic acids derived from the microbial sample.
- Hybridization ("Dips"): Chip is incubated with the target solution; specific binding reveals target presence/abundance.
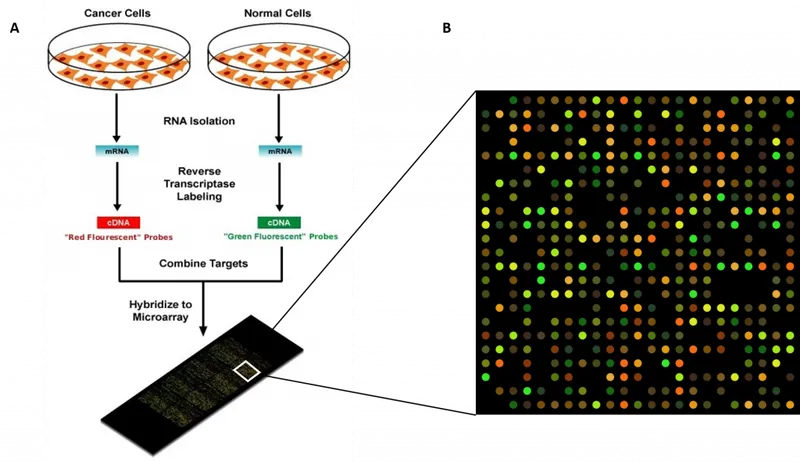
⭐ DNA microarrays can simultaneously detect multiple pathogens and their antibiotic resistance genes from a single clinical sample.
Microarray Workflow - Lab Bench Ballet
- Principle: Utilizes massively parallel hybridization on a chip. DNA probes bind labeled sample nucleic acids for simultaneous multi-target detection.
- Steps:
- Sample preparation: Isolate & purify microbial DNA/RNA (e.g., from patient specimen).
- Labeling: Enzymatically attach fluorescent dyes (e.g., Cy3, Cy5) to sample nucleic acids.
- Hybridization: Incubate labeled sample with microarray chip; specific probe-target binding occurs.
- Washing: Stringent washes remove unbound/weakly bound material, reducing background noise.
- Scanning: Laser excites dyes; a scanner detects and quantifies emitted fluorescence intensity.
- Data analysis: Software processes signals, normalizes data, and interprets patterns for pathogen ID or gene analysis.
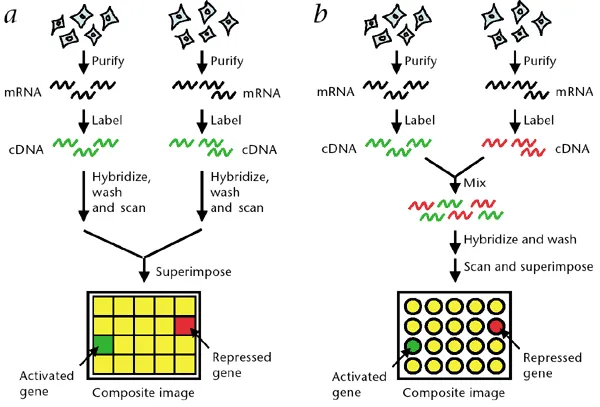
⭐ Microarrays enable high-throughput screening for multiple pathogens or AMR genes in one assay, aiding rapid diagnostics.
Diagnostic Applications - Bugs on Chips
- Multiplex Pathogen Detection:
- Simultaneously identifies diverse microbes (bacteria, viruses, fungi, parasites) from clinical samples (blood, sputum, CSF).
- Applications: Sepsis panels, respiratory virus panels, GI panels.
- Enables rapid species/strain level identification.
- Antimicrobial Resistance (AMR) Gene Detection:
- Identifies key resistance genes (e.g., $mecA$, $vanA$, $bla_{KPC}$, $bla_{NDM}$, ESBLs).
- Crucial for guiding targeted antibiotic therapy, combating resistance.
- Epidemiological Tool:
- Facilitates outbreak investigations & molecular epidemiology.
- Tracks pathogen transmission routes & evolution.
- Advantages: High throughput, speed, comprehensive screening from minimal sample.
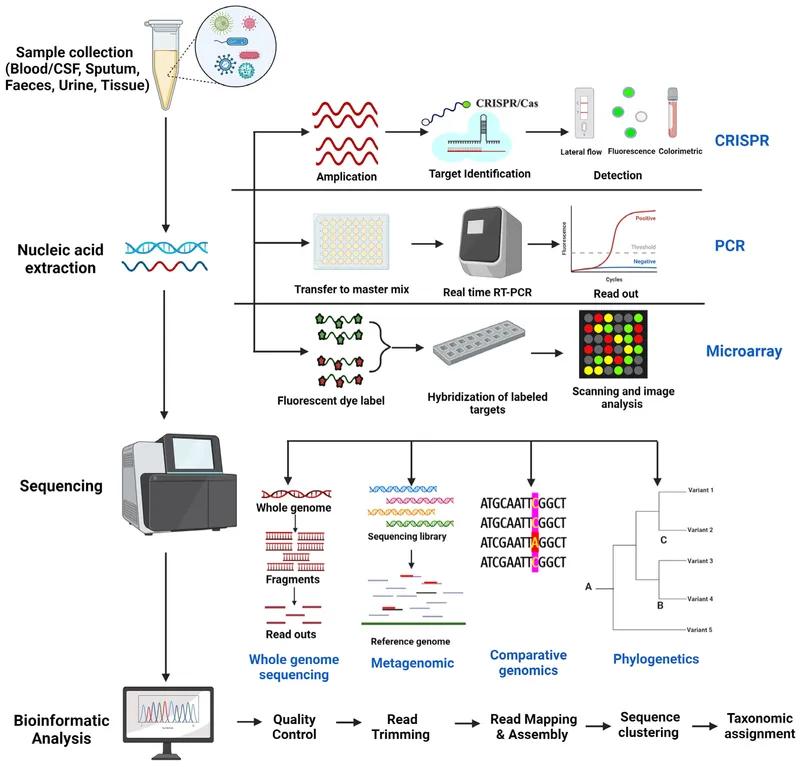
⭐ Microarrays allow for the detection of unculturable or slow-growing microorganisms, significantly reducing diagnostic turnaround time.
Clinical Utility & Future - Array of Hope?
- Clinical Utility:
- High-throughput: Simultaneous detection of multiple pathogens, virulence factors, & AMR genes.
- Rapid ID: Faster than culture for some organisms; detects unculturable microbes.
- Applications: Syndromic panels (respiratory, GI, sepsis), TB drug resistance (Line Probe Assays), viral genotyping.
- Limitations:
- Cost & complexity: Significant barriers in many Indian settings.
- Data interpretation can be challenging.
- Detects only pre-defined targets on the array.
- Future Prospects:
- Affordable Point-of-Care (POC) arrays.
- Integration with NGS for broader pathogen discovery.
- Enhanced epidemiological surveillance & outbreak response.
- Guiding personalized antimicrobial therapy.
⭐ Microarrays enable rapid, multiplex detection of antimicrobial resistance (AMR) genes, crucial for guiding timely and effective treatment in critical infections.
High‑Yield Points - ⚡ Biggest Takeaways
- Microarrays enable simultaneous detection of multiple microbial nucleic acids via hybridization.
- Critical for rapid pathogen identification, AMR gene screening, and outbreak analysis.
- Utilizes immobilized DNA probes (oligonucleotides or cDNA) on a chip.
- Offers high-throughput and multiplexing advantages for complex samples.
- Widely used for syndromic panels in sepsis, respiratory, and GI infections.
- Fluorescent signals typically indicate positive hybridization and target presence.
- Main limitations: cost and reliance on pre-existing sequence information for probe design.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

