Protein Folding Fundamentals - Nature's Origami
- Process: Polypeptide chain folds into a unique, biologically active 3D native conformation.
- Anfinsen's Dogma: Primary amino acid sequence contains all information for correct folding.
- Driving Forces:
- Hydrophobic effect (dominant): Nonpolar residues sequestered from water.
- Hydrogen bonds: Stabilize secondary (α-helices, β-sheets) and tertiary structures.
- Van der Waals forces: Weak, short-range attractions.
- Ionic bonds (salt bridges): Between charged groups.
- Disulfide bonds (-S-S-): Covalent; stabilize structure.
- Thermodynamics: Native state is thermodynamically most stable (lowest Gibbs free energy, $ΔG < 0$).
- Levinthal's Paradox: Proteins fold rapidly via specific pathways, not random conformational search.
- Folding Funnel: Energy landscape guiding protein towards native state.
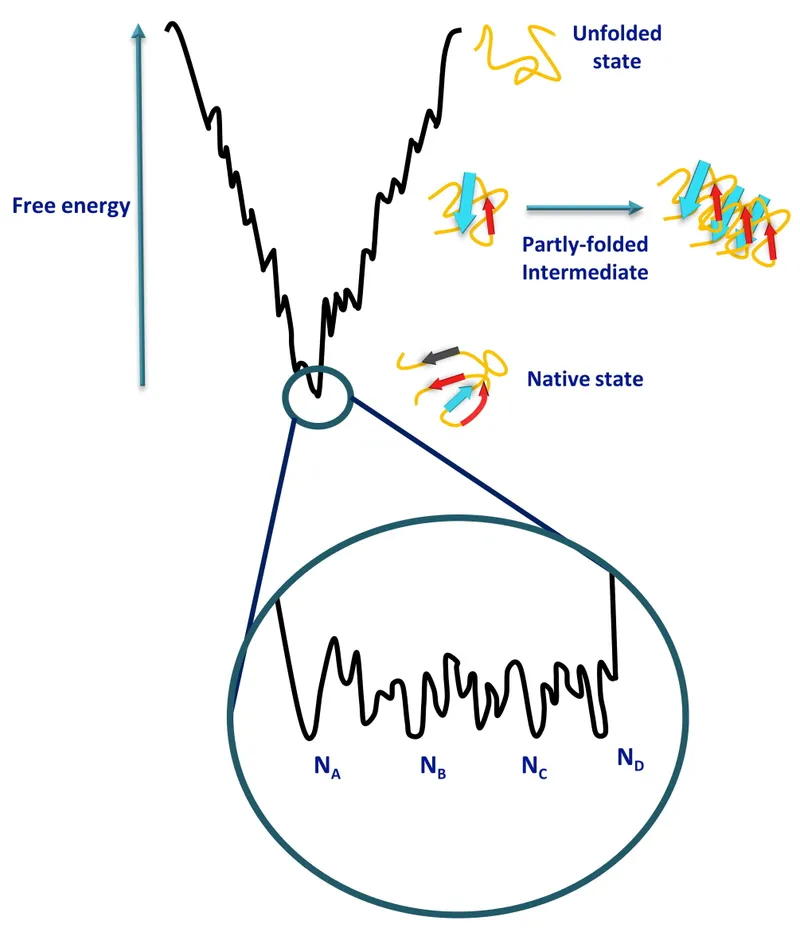
⭐ The hydrophobic effect, where nonpolar amino acid side chains cluster away from water, is the primary driving force for protein folding into its native conformation.
Molecular Chaperones & Mechanisms - Folding's Helping Hands
- Function: Proteins aiding correct folding of nascent/denatured polypeptides, prevent aggregation, assist translocation. Mostly ATP-dependent.
- Key Types & Actions:
- Hsp70 (e.g., DnaK in E.coli):
- Binds exposed hydrophobic patches on unfolded proteins.
- Acts co-translationally and post-translationally.
- Cycle: Hsp40 (DnaJ) aids substrate binding & ATP hydrolysis (tight grip); NEF (GrpE) promotes ADP/ATP exchange for release.
- 📌 Mnemonic: 70 is when you need help (early stage, versatile).
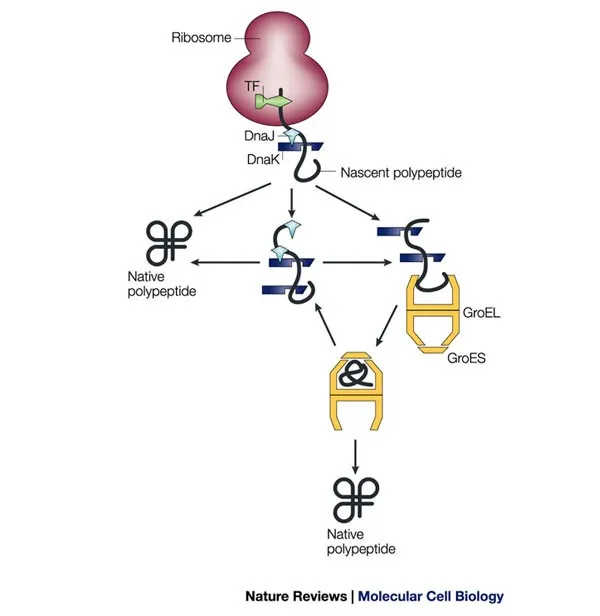
- Chaperonins (Hsp60 / GroEL-GroES complex in E.coli; TRiC/CCT in Eukaryotes):
- GroEL forms barrel; GroES acts as lid.
- Isolates unfolded proteins in cavity for undisturbed folding.
- ATP-dependent.
- 📌 Mnemonic: A 60-sided barrel (complex structure) for tough folding jobs.
- Hsp90:
- Late-stage folding/activation of specific "client" proteins (e.g., steroid receptors, signaling kinases).
- ATP-dependent.
- Calnexin & Calreticulin:
- ER lumen; lectin-like, bind N-glycans on glycoproteins. Ensure quality control.
- Hsp70 (e.g., DnaK in E.coli):
⭐ Hsp90 inhibitors (e.g., Geldanamycin, 17-AAG) are potent anti-cancer drug candidates, targeting the stability of numerous oncogenic client proteins.
- Pathology Link: Chaperone defects or overload implicated in protein misfolding diseases (e.g., Alzheimer's, Parkinson's, Cystic Fibrosis, Prion diseases).
Protein Misfolding & Diseases - Misfolded Mayhem
- Improper protein folding or misfolded protein accumulation causes cellular dysfunction and disease.
- Pathogenic Mechanisms:
- Loss of function: Degraded or non-functional misfolded protein (e.g., CFTR in Cystic Fibrosis).
- Gain of toxic function: Misfolded protein gains harmful activity (e.g., Huntington causing aggregation).
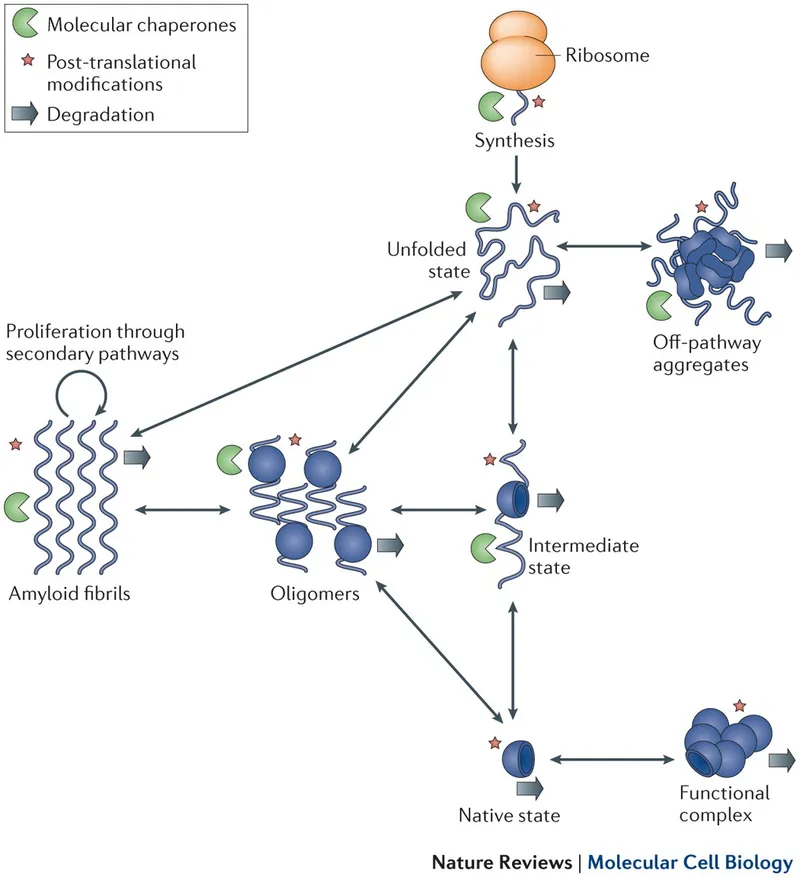
- Protein Aggregation & Amyloid Formation: Misfolded proteins form insoluble amyloid aggregates.
- Often β-sheet rich, proteolysis-resistant.
- Key Proteopathies (Misfolding Diseases):
- Alzheimer's Disease: Extracellular Amyloid-β (Aβ) plaques; intracellular Tau neurofibrillary tangles.
- Parkinson's Disease: α-synuclein aggregation into Lewy bodies.
- Prion Diseases (e.g., CJD, Kuru): Pathogenic $PrP^{Sc}$ induces misfolding of normal $PrP^{C}$.
- Cystic Fibrosis: CFTR misfolding (esp. $\Delta F508$ mutation) leads to premature degradation.
- Amyloidosis: Systemic or localized amyloid deposition.
⭐ Prion diseases are unique as the misfolded PrPSc protein can act as a template, inducing misfolding in normal PrPC proteins, making them transmissible.
High‑Yield Points - ⚡ Biggest Takeaways
- Protein folding yields the functional native 3D structure, determined by the amino acid sequence.
- Anfinsen's experiment proved primary structure dictates tertiary structure.
- Molecular chaperones (e.g., Hsp70, Hsp60/GroEL-GroES) aid correct folding and prevent aggregation.
- Misfolded proteins are linked to diseases like Alzheimer's, Parkinson's, and prion diseases.
- Levinthal's paradox suggests folding follows defined pathways, not random search.
- PDI (Protein Disulfide Isomerase) assists disulfide bonds; PPI (Peptidyl Prolyl Isomerase) aids proline isomerization.
- The ubiquitin-proteasome pathway degrades misfolded proteins.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

