Genetic Code Basics - Cracking Life's Cipher
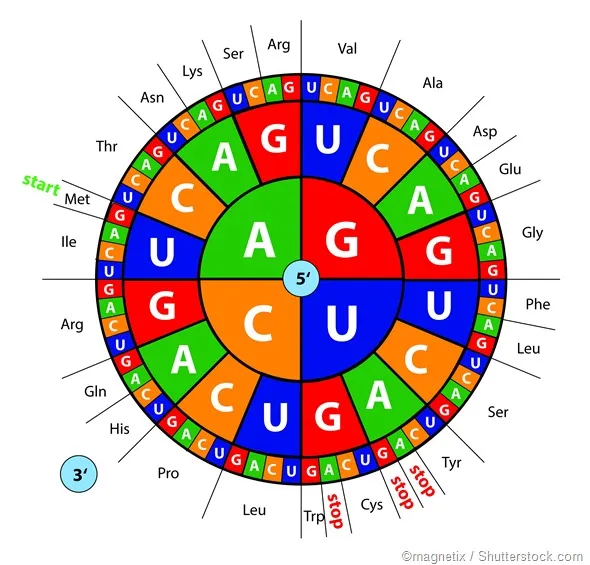
- Triplet Code: 3 nucleotide bases (codon) specify one amino acid.
- Codons: 64 total ($4^3$); 61 sense (amino acids), 3 stop (UAA, UAG, UGA).
- Start Codon: AUG (Methionine); initiates protein synthesis.
- Key Features:
- Unambiguous: One codon → specific amino acid.
- Degenerate: Multiple codons → same amino acid.
- Non-overlapping, commaless.
- Universal (mostly; mitochondrial exceptions).
⭐ Wobble Hypothesis: Flexible base pairing at 3rd codon position allows some tRNAs to recognize multiple codons, explaining degeneracy efficiently.
Code's Quirks - Rules of the Game
- Degenerate (Redundant): Multiple codons specify a single amino acid.
- E.g., Leucine has 6 codons.
- Only Methionine (AUG) and Tryptophan (UGG) have single codons.
- Wobble hypothesis (Crick): 3rd codon base has flexible pairing with anticodon's 1st base.
- Unambiguous: Each specific codon codes for only one amino acid. No codon specifies more than one.
- Non-overlapping: Codons are read sequentially in groups of three bases from a fixed starting point.
- Commaless: No punctuation or spacers between codons; continuous reading frame.
- Universal: Largely conserved across species, from bacteria to humans.
- Minor variations exist (e.g., mitochondrial DNA, Mycoplasma, some ciliates).
- Start Codon: AUG (Methionine). GUG can also initiate (Valine) in some cases.
- Stop Codons (Nonsense): UAA (Ochre), UAG (Amber), UGA (Opal). 📌 U Are Away, U Are Gone, U Go Away.
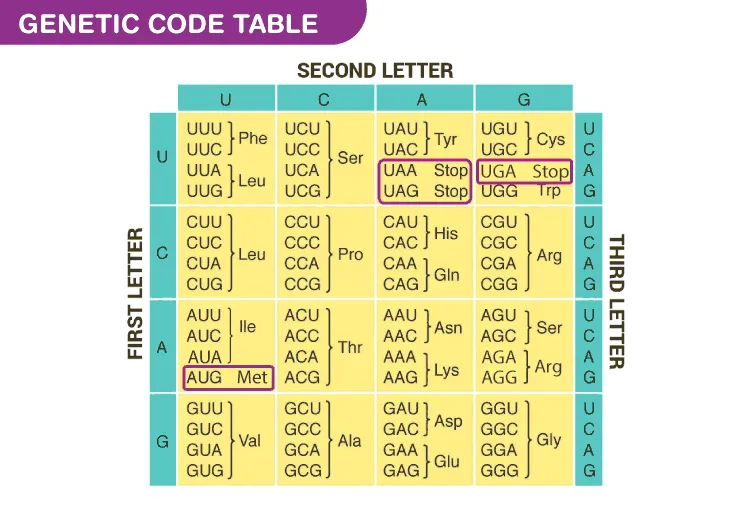
⭐ UGA, typically a stop codon, codes for Selenocysteine (the 21st amino acid) in specific contexts, requiring a downstream SECIS element (Selenocysteine Insertion Sequence).
Wobble & tRNA - Flexible Friends
- Wobble Hypothesis (Crick): Non-Watson-Crick base pairing at the codon's 3rd base (mRNA 3' end) and the anticodon's 1st base (tRNA 5' end).
- Allows one tRNA to recognize multiple codons, reducing the number of tRNAs needed.
- **Key Wobble Pairs (Anticodon base ↔ Codon base):
- G ↔ U or C
- U ↔ A or G
- I (Inosine) ↔ U, C, or A 📌 Mnemonic: "I See You All" (C, U, A)
- Significance: Fewer tRNAs required (e.g., humans use ~48 tRNAs for 61 sense codons).
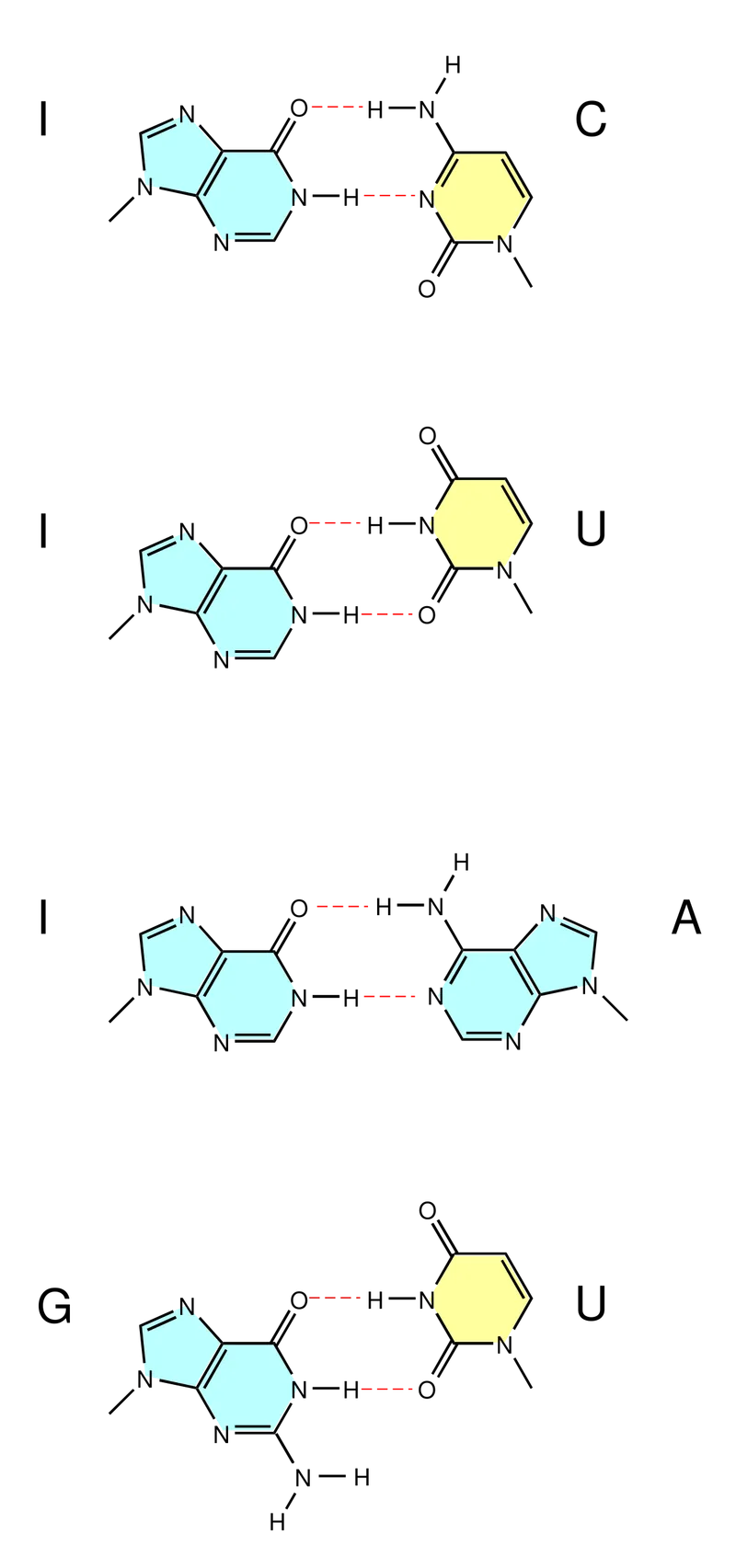
⭐ Inosine (I), formed by post-transcriptional deamination of adenosine in the tRNA anticodon, allows for the most extensive wobble pairing (recognizing U, C, and A).
Codon Bias & Blips - Code in Action
- Codon Usage Bias:
- Unequal use of synonymous codons; species-specific.
- Reflects tRNA availability; influences mRNA stability & translation rate.
- Key for optimizing recombinant protein expression.
- Genetic Code "Blips" (Mutations):
- Point Mutations:
- Silent: Different codon, same amino acid (wobble effect).
- Missense: Changes amino acid; may alter protein function.
- Nonsense: Forms STOP codon (UAA, UAG, UGA) → premature termination. 📌 Remember: "U Are Away, U Are Gone, U Go Away".
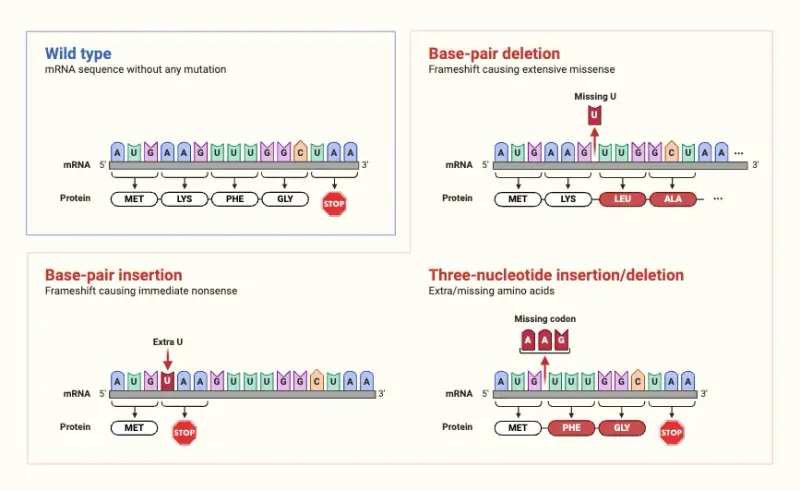
- Frameshift Mutations: Insertions/deletions (not multiples of 3) → altered reading frame, often creating premature stop codons downstream & non-functional protein.
- Point Mutations:
⭐ Sickle cell anemia is a classic example of a missense mutation (GAG to GUG in β-globin gene), leading to HbS.
High‑Yield Points - ⚡ Biggest Takeaways
- Genetic code: degenerate, unambiguous, non-overlapping, commaless, and nearly universal.
- Start codon: AUG (Methionine). Stop codons: UAA (Ochre), UAG (Amber), UGA (Opal).
- Wobble hypothesis: flexible base pairing at the 3rd codon position allows some tRNAs to recognize multiple codons.
- Mitochondrial DNA shows code variations: e.g., UGA as Tryptophan, AGA/AGG as stop codons.
- Codon usage bias: preferential use of certain synonymous codons varies between organisms and genes.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

