Single Nucleotide Polymorphisms - Tiny Changes, Big Impact
- Single Nucleotide Polymorphisms (SNPs): DNA sequence variations affecting a single nucleotide.
- Prevalence: Most common genetic variant; present in >1% of the population.
- Location: Found in coding (exons), non-coding (introns, regulatory), or intergenic regions.
- Coding region SNP effects:
- Synonymous (silent): Different codon, same amino acid.
- Non-synonymous: Results in amino acid change.
- Missense: Different amino acid.
- Nonsense: Premature stop codon.
- Coding region SNP effects:
- Significance: Disease susceptibility, pharmacogenomics (drug response), population markers, forensics. within a DNA strand compared to a reference sequence)
⭐ SNPs are crucial for Genome-Wide Association Studies (GWAS), linking genetic variants to diseases and traits.
SNP Types & Effects - Code Breakers
- Coding SNPs (cSNPs): Impact protein.
- Synonymous (Silent): No amino acid change; often benign.
- Non-synonymous:
- Missense: Alters one amino acid; variable effect.
- Nonsense: Premature STOP codon; truncated, non-functional protein.
- Non-coding SNPs: Affect gene activity/splicing.
- Regulatory: In promoters/enhancers; alter gene expression (↑/↓).
- Intronic: Within introns; can disrupt mRNA splicing.
- Intergenic: Between genes; may affect distant gene regulation.
Key Impacts:
- Disease risk (e.g., complex traits).
- Drug response variability (pharmacogenomics).
- Phenotypic differences.
⭐ > Non-synonymous SNPs are a major focus in identifying genetic causes of Mendelian diseases.
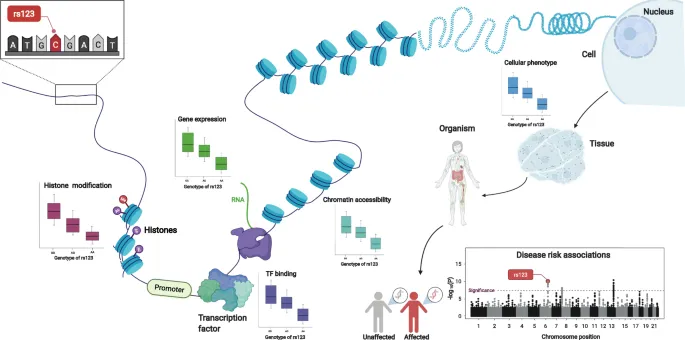
Finding SNPs - Genetic Detectives
SNP discovery identifies single base variations by comparing DNA sequences. Key steps involve sample preparation, data generation, and bioinformatic analysis.
- Primary Approaches:
- DNA Sequencing:
- Next-Generation Sequencing (NGS): WGS (Whole Genome) & WES (Whole Exome) for comprehensive, novel SNP discovery. Aligned to reference genome.
- Sanger Sequencing: Gold-standard for targeted SNP validation.
- Microarrays (SNP Chips):
- High-throughput genotyping of thousands to millions of known SNPs.
- Based on allele-specific oligonucleotide hybridization.
- PCR-Based Methods:
- For specific, known SNPs (e.g., RFLP, TaqMan assays, ARMS-PCR). Often used for validation or smaller scale studies.
- DNA Sequencing:
⭐ GWAS (Genome-Wide Association Studies) heavily rely on SNP microarrays to find associations between SNPs and traits/diseases in large cohorts.
SNPs in Medicine - Clinical Clues
- Disease Susceptibility: SNPs can ↑ or ↓ risk for common diseases (e.g., diabetes, heart disease, autoimmune disorders).
- APOE gene SNPs (ε2, ε3, ε4 alleles) and Alzheimer's disease risk.
- ε4 allele: ↑ risk, earlier onset.
- ε2 allele: protective effect.
- APOE gene SNPs (ε2, ε3, ε4 alleles) and Alzheimer's disease risk.
- Pharmacogenomics: SNPs influence drug efficacy and adverse drug reactions (ADRs).
- CYP2C19 SNPs: Clopidogrel metabolism (poor vs. extensive metabolizers).
- VKORC1 SNPs: Warfarin sensitivity, dose adjustments.
- TPMT SNPs: Azathioprine toxicity risk.
- HLA-B5701*: Abacavir hypersensitivity.
- Cancer: Somatic SNPs in tumors (drivers/passengers); germline SNPs for predisposition (e.g., BRCA1/2 variants, though often not single SNPs).

⭐ Warfarin Dosing: SNPs in CYP2C9 (drug metabolism) and VKORC1 (drug target) genes are critical for determining appropriate warfarin dosage, significantly impacting anticoagulation therapy and reducing bleeding or clotting risks. This is a frequently tested pharmacogenomic application.
- Monogenic Diseases: While many SNPs are common, rare SNPs can be pathogenic and cause Mendelian disorders (e.g., Cystic Fibrosis - CFTR gene mutations, some of which are SNPs).
- Forensic Science: SNP profiling for individual identification (though less variable than STRs).
High‑Yield Points - ⚡ Biggest Takeaways
- SNPs: Most common genetic variation; single base-pair difference.
- Must be present in >1% of the population.
- Occur in coding (exons) or non-coding (introns, regulatory) regions.
- Impact disease susceptibility, drug metabolism (pharmacogenomics), and phenotypic traits.
- Non-synonymous SNPs alter protein sequence; synonymous SNPs do not.
- Crucial for GWAS, linkage disequilibrium (LD) analysis, and haplotype mapping.
- Detected via DNA microarrays and Next-Generation Sequencing (NGS).
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

