PCR: Introduction - Chain Reaction Magic
- Polymerase Chain Reaction (PCR): A cornerstone in vitro technique for exponential amplification of specific DNA sequences.
- Mimics natural DNA replication, creating millions to billions of copies from a minute starting sample, enabling detailed analysis.
- "Chain reaction": Products of each cycle serve as templates for subsequent cycles, leading to rapid, exponential increase.
⭐ PCR was developed by Kary Mullis in 1983, for which he received the Nobel Prize in Chemistry in 1993 an exam-favourite fact!
PCR: Components - The Reaction Cocktail
Key reagents for DNA amplification. 📌 Mnemonic: TP-DAB (Template, Primers, dNTPs, Polymerase (DNA), Buffer).
| Component | Role & Key Properties |
|---|---|
| DNA Template | Source DNA with target sequence. |
| Primers (Fwd/Rev) | Define target; provide 3'-OH for synthesis. ~18-25 bp. |
| DNA Polymerase | Synthesizes DNA. Taq polymerase optimal ~72°C. |
| dNTPs | Building blocks (dATP, dGTP, dCTP, dTTP). |
| Buffer | Maintains optimal pH (e.g., Tris-HCl pH 8.3-8.8), contains KCl. |
| $Mg^{2+}$ ions | Cofactor ($MgCl_2$); critical for polymerase activity. Optimal: 1.5-2.5 mM. |
PCR: Steps - The Heat Dance
📌 DAE: Denature-Anneal-Extend. PCR automates DNA replication via repeated thermal cycles. Each cycle typically doubles the target DNA. Amplification formula: $2^n$ (where n = number of cycles).
- 1. Denaturation: Heat to ~94-98°C; separates double-stranded DNA (dsDNA) into single strands (ssDNA).
- 2. Annealing: Cool to ~50-65°C; allows primers to bind (anneal) to complementary sequences on ssDNA.
- 3. Extension: Heat to ~72°C (optimal for Taq polymerase); enzyme extends primers, synthesizing new DNA strands.
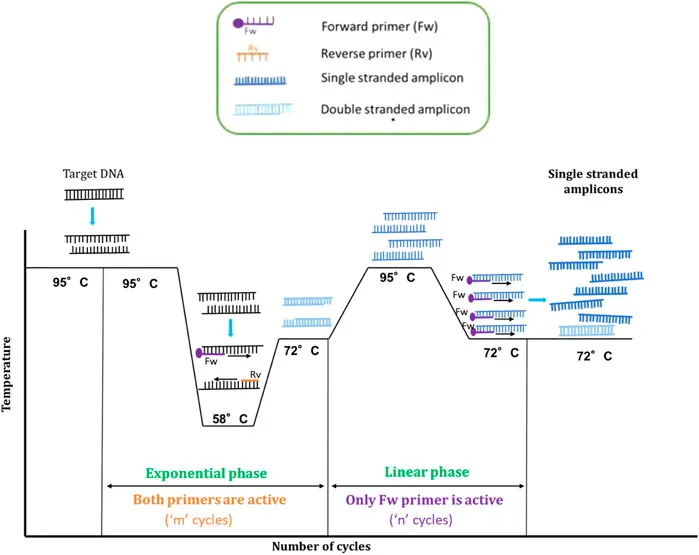
⭐ Annealing temperature is critical and depends on primer length and G-C content; too low = non-specific binding, too high = poor annealing.
PCR: Variants - Beyond The Basics
- Key PCR modifications enhance utility: 📌 Really Quick, Neat & Mighty! (RT, qPCR, Nested, Multiplex)
| Variant | Principle | Unique Application(s) |
|---|---|---|
| RT-PCR | RNA → cDNA (reverse transcriptase), then PCR. | RNA virus detection, gene expression (mRNA). |
| qPCR | Real-time fluorescence monitors DNA; $C_t$ value quantifies. | DNA/RNA quantification, viral load. |
| Nested PCR | Two PCRs: outer primers, then inner (nested) primers. | ↑ Specificity/sensitivity, low-abundance DNA. |
| Multiplex PCR | Simultaneous amplification of multiple targets, multiple primer sets. | Multi-pathogen ID, SNP genotyping. |
PCR: Applications & Pitfalls - Uses & Cautions
- Applications
- Diagnosis: Infectious (HIV, TB, COVID-19), genetic diseases (cystic fibrosis).
- Forensics: DNA fingerprinting, paternity testing.
- Research: Gene cloning, sequencing, site-directed mutagenesis.
- Prenatal diagnosis of inherited disorders.
- Advantages
- High sensitivity: Amplifies minute DNA.
- High specificity: With well-designed primers.
- Rapidity: Results within hours.
- Limitations & Cautions
- Contamination: High risk of false positives.
- Primer design: Crucial for success; non-specific binding.
- Taq polymerase errors: Lacks proofreading.
- Inhibitors: Present in clinical samples.
⭐ A major limitation of PCR is its susceptibility to contamination, leading to false-positive results.
High‑Yield Points - ⚡ Biggest Takeaways
- PCR is a rapid in-vitro DNA amplification technique, generating millions of copies.
- Key components: Taq polymerase (heat-stable), primers, dNTPs, and template DNA.
- Core steps per cycle: Denaturation (
95°C), Primer Annealing (50-65°C), Extension (~72°C).- Achieves exponential amplification (2^n) of the target DNA.
- Vital for diagnosing infections, genetic testing, forensics, and research.
- RT-PCR detects RNA by first converting it to cDNA using reverse transcriptase.
- qPCR (Real-Time PCR) allows quantification of DNA/RNA during amplification.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

