Secondary Structure Basics - The Local Folds
- Local, regular, repeating 3D arrangements of the polypeptide backbone.
- Stabilized mainly by hydrogen bonds between backbone C=O and N-H groups.
- Peptide Bond:
- Planar & rigid (partial double bond character).
- No rotation around C-N bond.
- Key Dihedral Angles:
- Phi ($\phi$): Rotation around N-C$\alpha$ bond.
- Psi ($\psi$): Rotation around C$\alpha$-C bond.
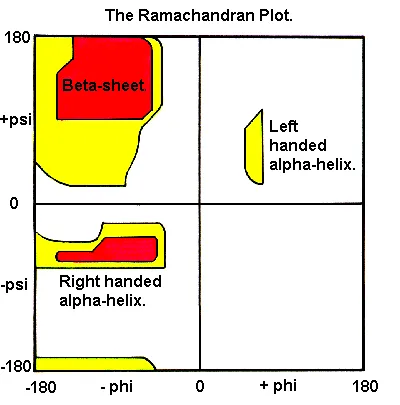
- Ramachandran Plot:
- Maps sterically allowed ($\phi$, $\psi$) combinations.
- Predicts stable conformations.
⭐ Hydrogen bonds between backbone amide (N-H) and carbonyl (C=O) groups are crucial for stabilizing secondary structures.
Alpha-Helix - Coiling Up Right
- Right-handed coiled polypeptide chain; most common regular secondary structure.
- Stabilized by intrachain H-bonds:
- Between carbonyl O (residue $n$) & amide H (residue $n+4$).
- Key Parameters:
- 3.6 residues per turn.
- Pitch (axial distance/turn): 5.4 Å.
- Rise per residue: 1.5 Å.
- Amino Acids:
- Formers: Met, Ala, Leu, Glu, Lys (📌 Mnemonic: MALEK).
- Breakers: Proline (causes kink, no N-H for H-bond), Glycine (too flexible).
⭐ Proline frequently disrupts alpha-helices because its ring structure creates a kink and it cannot form the necessary hydrogen bond.
Beta-Pleated Sheet - Folding Flat Out
- Composed of extended polypeptide segments called β-strands.
- Strands align laterally, forming a sheet; stabilized by inter-strand H-bonds (C=O···H-N).
- R-groups project alternately above and below the pleated sheet plane.
- Often rich in Glycine (Gly) & Alanine (Ala) due to small R-groups.
| Feature | Parallel β-Sheet | Anti-parallel β-Sheet |
|---|---|---|
| Strand Direction | Same (N→C | |
| H-bond Geometry | Bent, weaker | Linear, stronger |
| Relative Stability | Less stable | More stable |
⭐ Silk fibroin is predominantly composed of anti-parallel β-pleated sheets, contributing to its strength and flexibility.
Turns & Loops - Connecting The Dots
- Connect α-helices & β-sheets; change chain direction abruptly.
- β-Turns (Reverse Turns):
- Connect anti-parallel β-strands.
- 4 amino acid residues. H-bond: CO (residue $i$) & NH (residue $i+3$).
- Pro (pos 2) & Gly (pos 3) common. 📌 Pro-Gly "Turn Pair".
- Pro: fixed φ angle for turn.
- Gly: small, flexible.
- Types I & II common.
- γ-Turns:
- 3 amino acid residues. H-bond: CO (residue $i$) & NH (residue $i+2$).
- Loops:
- Larger, less regular connecting segments.
- Surface loops: interactions, recognition, active sites.
-
⭐ Pro & Gly: "helix breakers", but crucial "turn makers". oka
Motifs & Fibers - Building Blocks & Backbones
- Supersecondary Structures (Motifs): α-helix & β-sheet combinations.
- βαβ unit: α-helix links two parallel β-strands.
- β-hairpin: Antiparallel β-strands connected by a turn.
- Greek key: Four antiparallel β-strands forming a pattern.
- Fibrous Proteins: Elongated, structural roles; repetitive sequences.
- α-Keratin: Coiled-coil of two α-helices (hair, nails); Cys-rich (disulfide bonds).
- Silk Fibroin: Stacked antiparallel β-sheets (silk); Ala/Gly-rich; flexible.
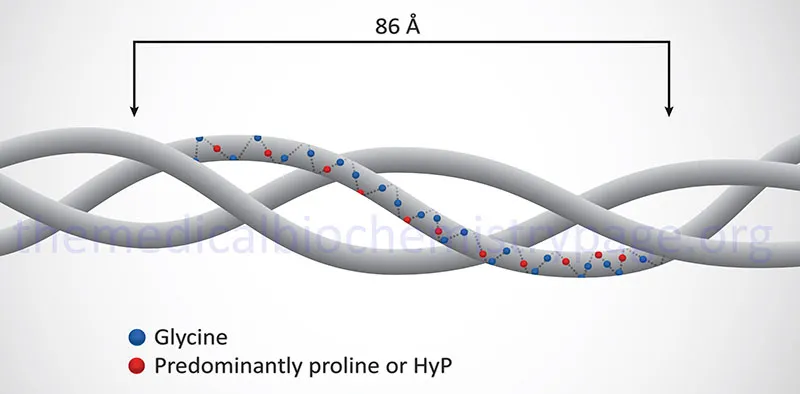
- Collagen: Unique triple helix (Gly-X-Y repeats; X=Pro, Y=Hyp). 📌 Glycine every 3rd residue.
⭐ Collagen: most abundant animal protein. Vitamin C needed for Pro/Lys hydroxylation, vital for stability.
High‑Yield Points - ⚡ Biggest Takeaways
- α-helix & β-pleated sheet are key secondary structures.
- Stabilized mainly by hydrogen bonds within the peptide backbone.
- α-helix: Right-handed coil, 3.6 residues/turn; features intrachain H-bonds.
- Proline (helix kink) & Glycine (too flexible) act as helix breakers.
- β-sheet: Comprises β-strands (parallel/antiparallel); H-bonds are often interchain.
- β-turns (reverse turns) connect antiparallel β-sheet strands, frequently containing Proline & Glycine.
- Ramachandran plot visualizes allowed φ and ψ backbone dihedral angles.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more


