Protein Domains and Motifs - Protein Lego Land
- Protein Domain:
- A stable, independently folding unit within a protein.
- Has a specific function (e.g., binding, catalysis).
- Fundamental unit of structure, function, and evolution.
- Protein Motif (Supersecondary Structure):
- A short, conserved sequence/structural arrangement.
- Often part of a domain; may not be stable or functional alone.
- Examples: β-α-β unit, Greek key, helix-loop-helix.
- Structural Hierarchy:
- Primary (Sequence) → Secondary (α-helix, β-sheet) → Motifs → Tertiary (Domains, 3D shape) → Quaternary.
- Domain vs. Motif:
- Domain: Independent functional & structural unit.
- Motif: Structural pattern; function often tied to its domain context.
- Importance:
- Modularity: Proteins are like "Lego" structures built from domains.
- Evolution: New functions via domain shuffling.
- Prediction: Domains/motifs aid in predicting protein roles.
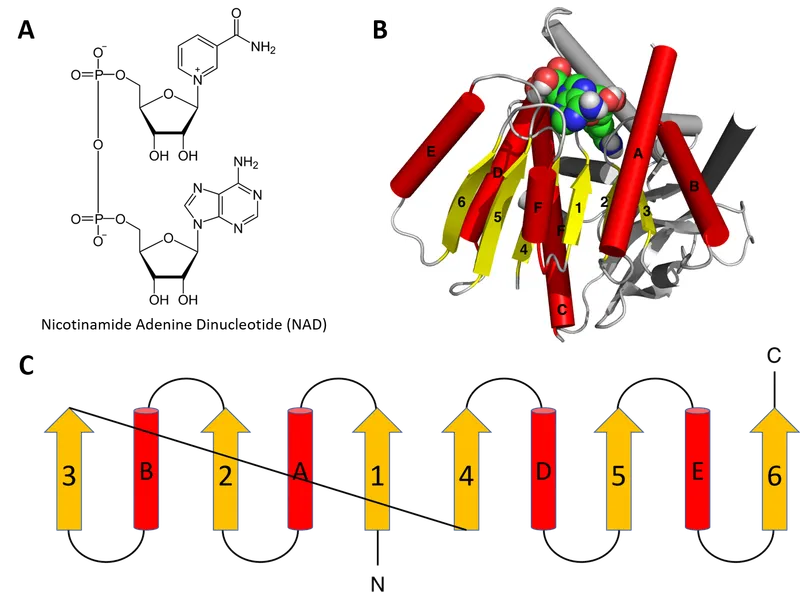
⭐ Protein domains are often considered the fundamental units of protein evolution.
Protein Domains and Motifs - Functional Hotspots
Protein domains are conserved, independently folding functional units within a protein. Motifs are shorter, recurring sequence patterns. These are crucial for protein interactions and cellular signaling.
| Domain Name | Recognized Ligand/Site | Key Function(s) | Clinical Relevance Example |
|---|---|---|---|
| SH2 (Src Homology 2) | Phospho-Tyrosine (pY) | Signal transduction, protein-protein interaction | Chronic Myeloid Leukemia (BCR-ABL signaling) |
| SH3 (Src Homology 3) | Proline-rich sequences | Signal transduction, cytoskeletal organization | Adapter proteins in signaling (e.g., Grb2) |
| Pleckstrin Homology (PH) | Phosphoinositides (e.g., PIP3) | Membrane targeting, signal transduction | Akt/PKB signaling in cancer |
| Zinc Finger | DNA/RNA | Gene transcription, DNA repair, RNA binding | Transcription factor defects (e.g., steroid hormone receptors) |
| Kinase Domain | ATP, substrate protein | Catalyzes phosphorylation, signal amplification | Kinase inhibitors in cancer therapy (e.g., Imatinib) |
⭐ Many signaling proteins utilize SH2 domains to interact with activated receptor tyrosine kinases.
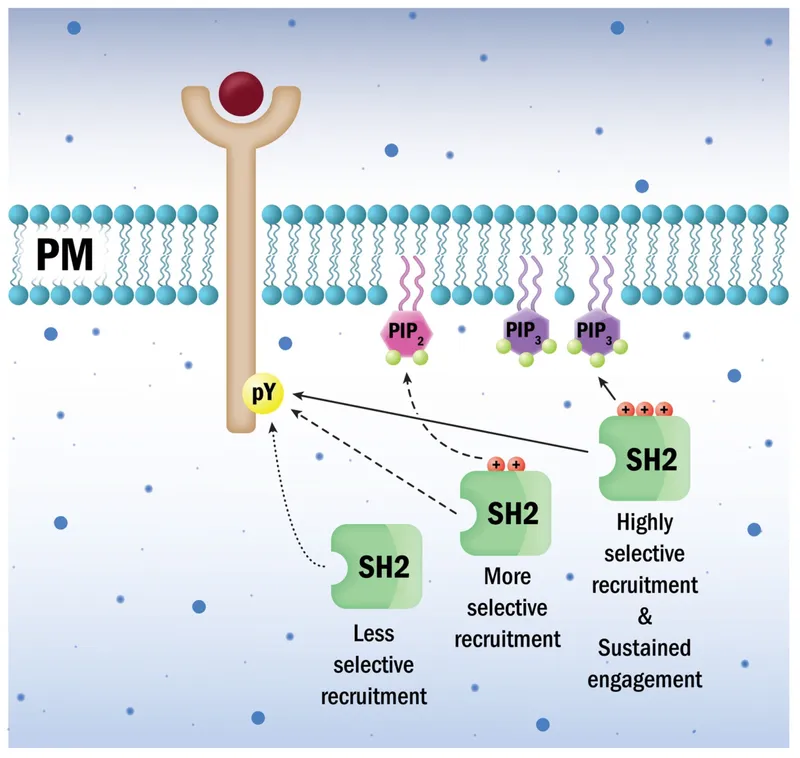
Protein Domains and Motifs - Pattern Play
- Motifs (Supersecondary Structures): Small, conserved structural units built from specific arrangements of secondary structure elements ($\alpha$-helices, $\beta$-strands, and turns). They are smaller than domains and often represent functional sites or form part of a larger domain.
| Motif Name | Structural Description | Common Role(s) | Example Protein(s) |
|---|---|---|---|
| Helix-turn-helix | Two $\alpha$-helices connected by a short amino acid sequence (turn) | DNA binding, gene regulation | Lac repressor, homeodomain proteins |
| Leucine zipper | Two amphipathic $\alpha$-helices with leucine at every 7th residue, forming a coiled-coil | Dimerization of transcription factors, DNA binding | c-Fos/c-Jun, CREB |
| Zinc finger | One or more Zn²⁺ ions coordinate with Cys/His residues, stabilizing a small fold | DNA/RNA binding, protein-protein interactions | TFIIIA, GATA factors, steroid receptors |
| EF hand | A helix-loop-helix structure; the loop binds Ca²⁺ ions | Calcium sensing and binding | Calmodulin, parvalbumin, troponin C |
| Beta-alpha-beta unit | Two parallel $\beta$-strands linked by an intervening $\alpha$-helix | Forms core of many nucleotide-binding domains, enzymes | Rossmann fold, dehydrogenases, kinases |
⭐ The helix-turn-helix motif is a common DNA-binding motif found in many prokaryotic and eukaryotic transcription factors.
High‑Yield Points - ⚡ Biggest Takeaways
- Protein domains are stable, independently folding globular units within a polypeptide, often associated with a specific function.
- Motifs (or supersecondary structures) are short, conserved sequence patterns or structural arrangements, like the helix-turn-helix or beta-alpha-beta unit.
- A single domain can comprise multiple motifs; motifs themselves typically do not fold independently.
- Modular evolution: Proteins often evolve by shuffling or combining existing domains.
- Key examples: Zinc finger (DNA binding), Leucine zipper (dimerization), EF hand (Ca²⁺ binding).
- Mutations within critical domains can impair protein function, leading to disease states (e.g., kinase domain mutations in cancer).
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

