RNA Structure Basics - Sugar, Base, Chain!
- RNA: Typically single-stranded polymer of ribonucleotides.
- Linkage: Covalent $3'-5'$ phosphodiester bonds form the backbone.
- Ribonucleotide = Ribose sugar + Phosphate group + Nitrogenous Base.
- Ribose: Pentose sugar with a reactive $2'-OH$ group (DNA's deoxyribose has $2'-H$).
- Bases:
- Purines: Adenine (A), Guanine (G).
- Pyrimidines: Cytosine (C), Uracil (U). 📌 RNA uses Uracil (U); DNA uses Thymine (T).
- Chain Polarity: Defined by $5'$ (phosphate) and $3'$ (hydroxyl) ends.

DNA vs. RNA
| Feature | RNA | DNA |
|---|---|---|
| Sugar | Ribose (with $2'-OH$) | Deoxyribose (lacks $2'-OH$, has $2'-H$) |
| Bases | A, G, C, U | A, G, C, T |
| Strands | Usually Single | Usually Double (helix) |
| Stability | Less stable (due to $2'-OH$) | More stable |
| Function | Protein synthesis, gene regulation, enzyme | Genetic information storage |
Major RNA Players - The RNA Trio
Three major RNA types are central to protein synthesis, each with distinct structures and functions:
| RNA Type | Full Name | % of Total RNA | Key Structural Features | Specific Functions | Site of Action | 📌 Mnemonic |
|---|---|---|---|---|---|---|
| mRNA | Messenger RNA | ~5% | Linear; 5' cap (7-methylG); 3' poly-A tail (euk); codons. | Carries genetic code from DNA to ribosome; template for protein synthesis. | Cytoplasm | 📌 mRNA: Messenger for codons |
| tRNA | Transfer RNA | ~15% | Cloverleaf; anticodon loop (reads mRNA); D & TΨC loops; acceptor stem (3'-CCA binds amino acid). | Transports specific amino acid to ribosome; matches anticodon to mRNA codon. | Cytoplasm | 📌 tRNA: Transports amino acids |
| rRNA | Ribosomal RNA | ~80% | Core of ribosome subunits (Prok: 16S, 23S, 5S; Euk: 18S, 28S, 5.8S, 5S); globular. | Structural backbone of ribosomes; catalytic (ribozyme: 23S/28S rRNA forms peptide bonds). | Cytoplasm | 📌 rRNA: Ribosomal factory |
⭐ Wobble Hypothesis: The third base of an mRNA codon can form non-canonical pairs with the first base of a tRNA anticodon. This allows a single tRNA to recognize multiple codons, optimizing translation.
Other Functional RNAs - RNA's Special Forces
- snRNA (Small nuclear RNA):
- Function: Pre-mRNA splicing (spliceosome component).
- Location: Nucleus.
- snoRNA (Small nucleolar RNA):
- Function: rRNA modification (methylation, pseudouridylation).
- Location: Nucleolus.
- miRNA (MicroRNA):
- Function: Post-transcriptional gene silencing (mRNA degradation/repression).
- Location: Cytoplasm.
- siRNA (Small interfering RNA):
- Function: RNA interference (gene silencing, viral defense). Often exogenous.
- Location: Cytoplasm.
- lncRNA (Long non-coding RNA):
- Function: Diverse gene regulation (e.g., X-chromosome inactivation). >200 nt.
- Location: Nucleus, cytoplasm.
- piRNA (Piwi-interacting RNA):
- Function: Transposon silencing in germline cells.
- Location: Germ cells (with Piwi proteins).
⭐ RNA interference (RNAi) mediated by siRNAs and miRNAs is a key mechanism for gene regulation and defense against viral infections.
RNA Folding Fun - Twists and Turns
RNA folds into complex 3D structures via:
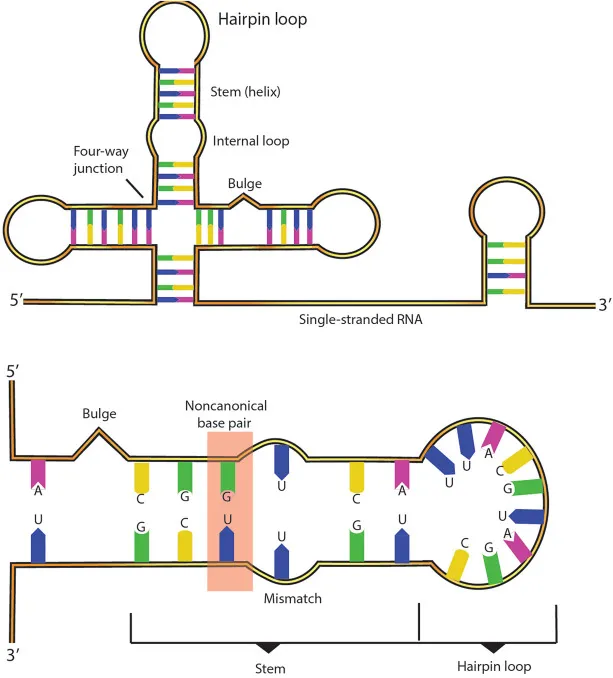
- Stem-loop (Hairpin): Intrastrand base pairing forms a duplex stem and an unpaired loop.
- Internal loop: Unpaired bases between two helical regions.
- Bulge: Unpaired nucleotide(s) on one side of a helix.
- Junctions: Intersection of multiple helical segments.
- Pseudoknot: Loop bases pair with a distant single-stranded region.
⭐ Pseudoknots are crucial for telomerase RNA and some viral RNAs.
- Non-canonical base pairs: (e.g., G-U wobble, Hoogsteen) stabilize tertiary structures.
High‑Yield Points - ⚡ Biggest Takeaways
- RNA: Typically single-stranded; contains ribose sugar and Uracil (U) instead of Thymine.
- mRNA: Carries genetic code from DNA to ribosome for protein synthesis.
- tRNA: Cloverleaf structure with an anticodon loop; transports specific amino acids.
- rRNA: Most abundant RNA; structural and catalytic (ribozyme) component of ribosomes.
- hnRNA: Eukaryotic precursor of mRNA; undergoes splicing, capping, and polyadenylation.
- Small RNAs: snRNA (involved in splicing), miRNA/siRNA (gene silencing).
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

