PTMs Intro - RNA's Makeover Magic
- Post-Transcriptional Modifications (PTMs): Covalent chemical alterations of RNA molecules (pre-mRNA, pre-tRNA, pre-rRNA) following transcription.
- Significance: Essential for RNA maturation, stability, transport, and function (e.g., translation, splicing).
- Location:
- Eukaryotes: Primarily nucleus (mRNA, tRNA, rRNA processing). Some tRNA mods in cytoplasm.
- Prokaryotes: Cytoplasm.
- Key RNA types undergoing PTMs: mRNA, tRNA, rRNA.
⭐ PTMs, like capping and polyadenylation in mRNA, are critical for its export from the nucleus and protection against exonucleases.
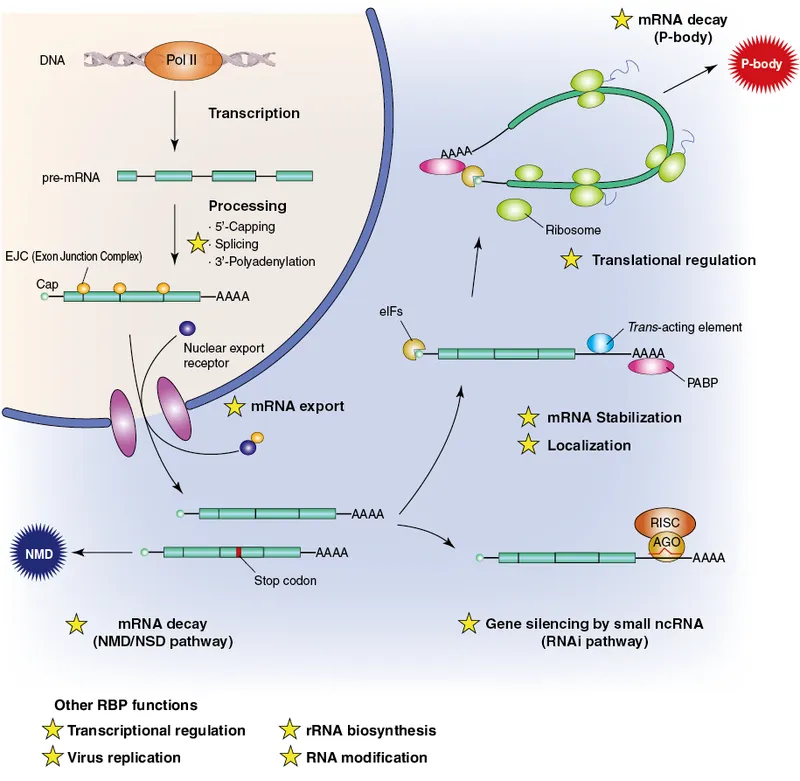
mRNA Capping & Tailing - mRNA's Hats & Tails
- 5' Capping: Addition of $m^7G$ (7-methylguanosine) to mRNA 5' end.
- Unusual 5'-5' triphosphate linkage.
- Occurs co-transcriptionally.
- Functions: Protects from 5' exonucleases, aids nuclear export, promotes translation initiation.
- 3' Polyadenylation (Tailing): Addition of ~50-250 adenine residues (poly-A tail) to mRNA 3' end.
- Signal: AAUAAA sequence.
- Enzyme: Poly(A) polymerase (template-independent).
- Functions: Protects from 3' exonucleases, aids nuclear export, enhances translation.
- 📌 Mnemonic: "Hats (caps) and Tails keep mRNA stable and sailing!"
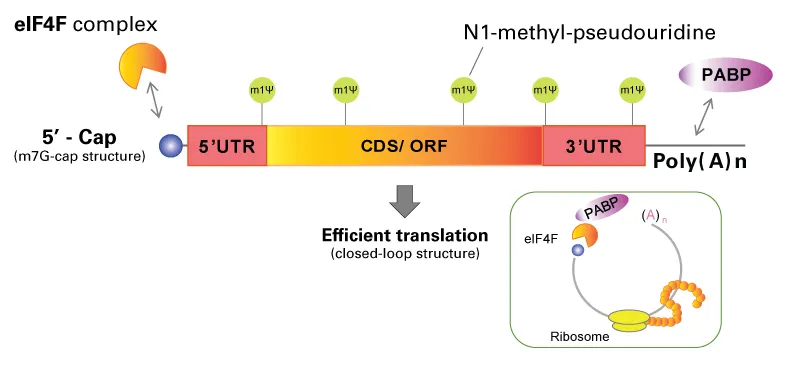
⭐ Histone mRNAs are a notable exception; they lack poly-A tails and instead have a 3' stem-loop structure.
mRNA Splicing - Splicing's Snip & Stitch
- Core Function: Nuclear process removing introns (non-coding) and ligating exons (coding) from pre-mRNA.
- Spliceosome: Large RNA-protein machine with snRNPs (U1, U2, U4, U5, U6).
- Consensus Sequences:
- 5' Splice Site (Donor): GU
- 3' Splice Site (Acceptor): AG
- Branch Point: Adenine (A) in intron near 3' site.
- 📌 Mnemonic: "Goo-Agro" for GU-AG intron boundary rule.
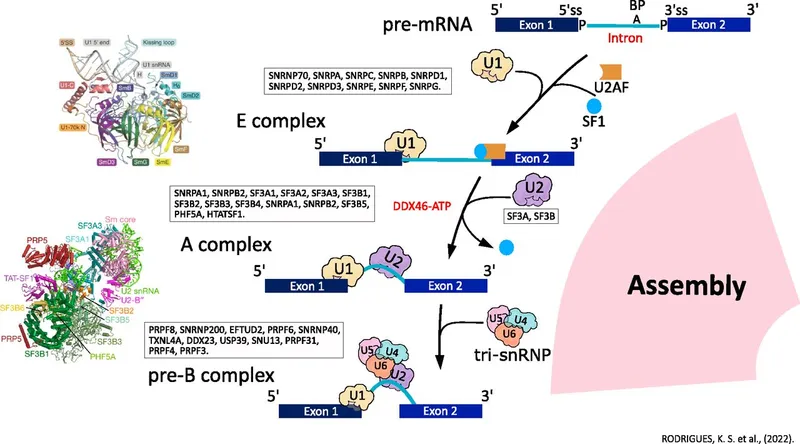
- Mechanism: Two transesterification steps:
- 1st: 2'-OH of branch point A attacks 5' splice site → lariat.
- 2nd: Free 3'-OH of 5' exon attacks 3' splice site → exons joined, lariat released.
- Alternative Splicing: One gene → multiple mRNA/protein isoforms; ↑ proteomic diversity.
⭐ Splicing errors contribute to ~15% of all human genetic diseases, like some β-thalassemias.
tRNA & rRNA Mods - RNA's Other Tweaks
- tRNA: Extensively modified (~25% bases) for structure, stability, codon recognition.
- Key Modified Bases:
Base Sym Significance Dihydrouridine $D$ D-loop, ↑ flexibility Pseudouridine $\Psi$ T$\Psi$C loop, stabilizes structure Ribothymidine $T$ T$\Psi$C loop, ribosome binding Inosine $I$ Wobble pairing (anticodon) - CCA Addition: Post-transcriptional at 3' end (tRNA nucleotidyltransferase); for amino acid attachment. 📌 "Can Carry Amino acids".
- Splicing of introns (some eukaryotic tRNAs).
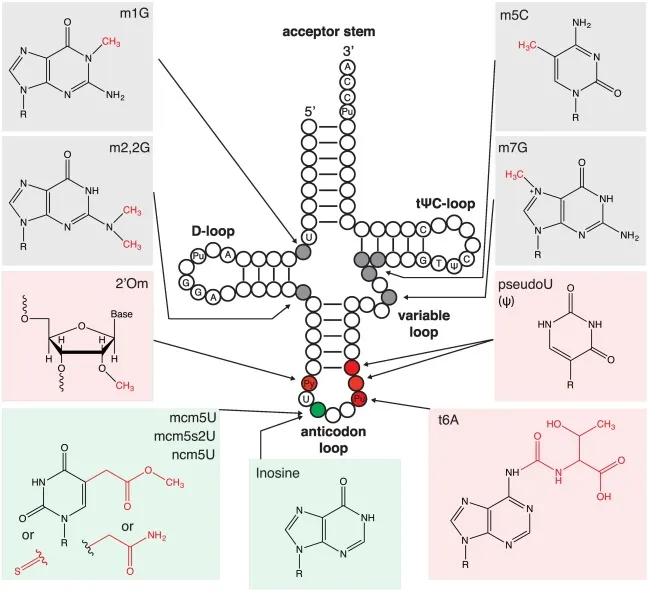
- Key Modified Bases:
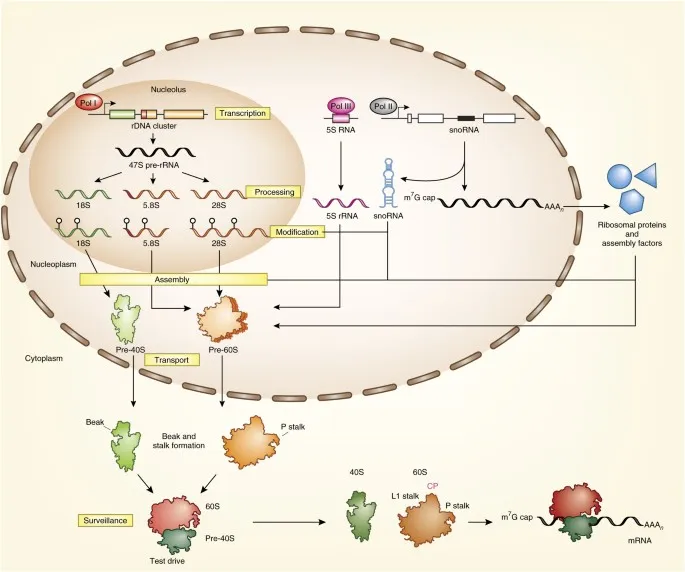
- rRNA: Processed from pre-rRNA (e.g., 45S → 18S, 5.8S, 28S in eukaryotes; 5S rRNA separate).
- Key Mods: Methylation, pseudouridylation; guide folding, nuclease protection.
- Significance: Crucial for ribosome assembly, structural integrity & catalytic activity.
⭐ The CCA sequence at the 3'-end of tRNA is added post-transcriptionally (not DNA-encoded) and is vital for aminoacylation.
High-Yield Points - ⚡ Biggest Takeaways
- 5' Capping: 7-methylguanosine (7-mG) cap addition; vital for mRNA stability & translation initiation.
- 3' Polyadenylation: Poly(A) tail addition by Poly(A) polymerase; boosts mRNA stability & translation.
- Splicing: Intron removal, exon joining by spliceosomes (snRNPs); a key nuclear process.
- Alternative Splicing: Generates protein isoforms from a single gene, expanding proteome diversity.
- RNA Editing: Post-transcriptional nucleotide changes; e.g., A-to-I (ADARs), C-to-U (ApoB mRNA).
- tRNA Modifications: Extensive base changes (e.g., pseudouridine) essential for tRNA structure and function.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

