Nucleic Acid Biochemistry

On this page
🧬 The Molecular Blueprint: Nucleic Acid Architecture Mastery
Every cell in your body stores, transmits, and executes genetic instructions through nucleic acids-molecules that determine everything from your eye color to your susceptibility to disease. You'll master the architecture of DNA and RNA, trace how genetic information flows from gene to protein, and develop the pattern recognition skills to identify pathological changes at the molecular level. By integrating structural knowledge with functional networks and clinical applications, you'll gain the diagnostic and therapeutic reasoning essential for interpreting genetic tests, understanding drug mechanisms, and recognizing how nucleic acid dysfunction drives human disease.
📌 Remember: DNA-RNA-Protein = Store-Transfer-Express - DNA stores genetic information with 99.9% fidelity, RNA transfers it with 85% efficiency, and proteins express it with 95% functional accuracy
The nucleic acid system operates through four fundamental levels: molecular structure (nucleotide composition), polymer organization (strand formation), three-dimensional architecture (helical geometry), and functional integration (information flow). Each level builds upon precise chemical interactions that determine genetic stability, replication fidelity, and expression control.
-
Structural Foundation
- Nucleotide components: phosphate + sugar + base = functional unit
- Sugar variations: ribose (RNA) vs deoxyribose (DNA) - single hydroxyl difference
- Ribose: 2'-OH present → increased reactivity, shorter half-life
- Deoxyribose: 2'-OH absent → enhanced stability, long-term storage
- Phosphodiester bonds: 3'-5' linkage with -7.3 kcal/mol bond energy
-
Base Pairing Architecture
- Purines (A, G): double-ring structures, larger molecular size
- Pyrimidines (C, T, U): single-ring structures, smaller molecular size
- Watson-Crick pairs: A-T (2 H-bonds), G-C (3 H-bonds)
- GC content correlation: higher GC = higher melting temperature
- Chargaff's ratios: A=T, G=C in double-stranded DNA
⭐ Clinical Pearl: GC-rich regions (>60% GC) create replication stress and mutation hotspots - explains why CpG islands are frequent sites of oncogenic mutations in 70% of human cancers
| Component | DNA | RNA | Clinical Significance | Stability | Function |
|---|---|---|---|---|---|
| Sugar | Deoxyribose | Ribose | DNA stable for forensics | High | Long-term storage |
| Bases | A,T,G,C | A,U,G,C | RNA degradation in samples | Low | Dynamic processing |
| Strands | Double | Single | PCR amplification design | Variable | Information transfer |
| Location | Nucleus | Cytoplasm | Biopsy interpretation | Protected | Active synthesis |
| Half-life | Years | Hours-Days | Sample handling protocols | Extended | Rapid turnover |
The molecular precision of nucleic acids enables error rates below 1 in 10^9 during replication, achieved through complementary base pairing (thermodynamic stability), proofreading mechanisms (3'-5' exonuclease activity), and mismatch repair systems (post-replication correction). This extraordinary fidelity makes genetic inheritance possible while allowing controlled mutation rates for evolutionary adaptation.
Connect this structural foundation through helical geometry principles to understand how three-dimensional architecture determines functional capacity and regulatory mechanisms.
🧬 The Molecular Blueprint: Nucleic Acid Architecture Mastery
⚡ The Information Highway: Nucleic Acid Functional Networks

The functional networks of nucleic acids operate through three integrated systems: information storage (genomic DNA organization), information transfer (transcriptional and translational machinery), and information regulation (epigenetic and post-transcriptional control). Each system maintains quality control checkpoints that ensure >99% accuracy in genetic information processing.
📌 Remember: TRIM = Transcription-Replication-Integration-Maintenance - the four core processes that maintain genetic information with error rates below 1 in 10^8 base pairs per cell division
-
Information Storage Architecture
- Genomic organization: 3.2 billion base pairs in human diploid genome
- Coding sequences: ~2% of genome (protein-coding genes)
- Exons: average 150 bp per coding sequence
- Introns: average 3,400 bp per non-coding sequence
- Regulatory regions: ~8% of genome (promoters, enhancers, silencers)
- Chromatin packaging: 147 bp DNA per nucleosome core particle
-
Information Transfer Mechanisms
- Transcription rates: 20-50 nucleotides/second for RNA polymerase II
- Translation rates: 15-20 amino acids/second for ribosomal synthesis
- mRNA half-life: 2-10 hours for most transcripts
- Protein half-life: 1-20 days for most cellular proteins
- Processing efficiency: >95% for constitutive exons, 60-90% for alternative exons
⭐ Clinical Pearl: Alternative splicing affects >90% of human genes, creating protein isoform diversity that explains how ~20,000 genes generate >100,000 different proteins - critical for understanding tissue-specific disease manifestations

| Process | Rate | Fidelity | Energy Cost | Clinical Relevance | Regulation |
|---|---|---|---|---|---|
| Replication | 50 bp/sec | 1 error/10^9 | 2 ATP/bp | Mutation analysis | Cell cycle |
| Transcription | 40 nt/sec | 1 error/10^4 | 2 ATP/nt | Expression profiling | Promoter activity |
| Translation | 15 aa/sec | 1 error/10^4 | 4 ATP/aa | Protein diagnostics | Ribosome availability |
| Repair | Variable | 99% correction | 10+ ATP/event | Cancer predisposition | Damage sensors |
| Splicing | 1-5 min/intron | 99.5% accuracy | 2 ATP/bond | Splice site mutations | SR proteins |
The information networks maintain homeostatic balance through feedback loops (negative regulation dominates), feedforward control (anticipatory responses), and checkpoint mechanisms (quality assurance). These control systems enable rapid adaptation (minutes to hours) while maintaining long-term stability (years to decades) of genetic information.
📌 Remember: PACE = Processing-Accuracy-Control-Efficiency - the four parameters that determine information network performance and predict clinical outcomes in genetic disorders and therapeutic responses
Connect this information processing framework through regulatory mechanism analysis to understand how control systems determine gene expression patterns and therapeutic targets.
⚡ The Information Highway: Nucleic Acid Functional Networks
🎯 The Diagnostic Decoder: Pattern Recognition in Nucleic Acid Analysis
Pattern recognition in nucleic acid analysis operates through four diagnostic frameworks: structural pattern analysis (sequence motifs and modifications), functional pattern correlation (expression signatures and pathway activation), temporal pattern tracking (dynamic changes and progression markers), and comparative pattern discrimination (normal vs pathological variants).
📌 Remember: SPOT = Sequence-Pattern-Output-Timing - the systematic approach for nucleic acid pattern recognition that achieves >90% diagnostic accuracy when applied consistently to molecular diagnostic interpretation
-
Sequence Pattern Recognition Framework
- Consensus sequences: TATA box (TATAAA at -25 to -30 bp), Kozak sequence (GCCRCCATGG)
- Splice sites: GT-AG (>99% of introns), AT-AC (<1% rare variants)
- Branch point: adenosine at -18 to -40 bp from 3' splice site
- Polypyrimidine tract: 10-15 consecutive pyrimidines upstream of AG
- Splice enhancers: ESE motifs (6-8 nucleotides) in exonic sequences
- Regulatory motifs: CpG islands (>50% GC, >60% CpG observed/expected)
-
Expression Pattern Correlation
- Housekeeping genes: coefficient of variation <20% across tissues
- Tissue-specific genes: >10-fold expression difference between tissues
- Developmental markers: temporal expression windows (hours to days)
- Stress response genes: 2-100 fold induction within 30 minutes
- Circadian genes: 24-hour oscillation with 2-5 fold amplitude
⭐ Clinical Pearl: CpG methylation patterns predict therapeutic response in >80% of cancers - hypermethylated promoters (>70% methylation) correlate with drug resistance, while hypomethylated regions (<30% methylation) indicate treatment sensitivity
| Pattern Type | Recognition Threshold | Clinical Sensitivity | Specificity | Diagnostic Window | Therapeutic Impact |
|---|---|---|---|---|---|
| Point mutations | >5% variant frequency | 85-95% | 99%+ | Single timepoint | Targeted therapy |
| Copy number | >1.5-fold change | 90-98% | 95-99% | Stable over time | Dosage adjustment |
| Methylation | >20% difference | 70-85% | 85-95% | Dynamic changes | Epigenetic drugs |
| Expression | >2-fold change | 80-90% | 70-85% | Temporal patterns | Pathway inhibitors |
| Splicing | >10% isoform shift | 75-90% | 90-95% | Tissue-dependent | Splice modulators |
- Cell cycle markers: G1/S (cyclin E peak), G2/M (cyclin B peak)
- Apoptosis cascade: caspase activation within 2-6 hours
- Early markers: phosphatidylserine externalization (30 minutes)
- Late markers: DNA fragmentation (2-4 hours)
- Recovery markers: anti-apoptotic protein upregulation (6-12 hours)
💡 Master This: Pattern integration across multiple molecular levels increases diagnostic accuracy from 70-80% (single marker) to >95% (multi-marker panels) - explains why comprehensive genomic profiling outperforms single-gene testing in cancer diagnosis and treatment selection
📌 Remember: WAVE = When-Amplitude-Variance-Expression - the four temporal parameters that characterize dynamic nucleic acid patterns and predict disease progression and treatment response timing
Connect this pattern recognition framework through comparative analysis methods to understand how systematic discrimination enables precise molecular diagnosis and personalized therapeutic strategies.
🎯 The Diagnostic Decoder: Pattern Recognition in Nucleic Acid Analysis
🔬 The Molecular Discriminator: Systematic Nucleic Acid Differentiation
📌 Remember: QUAD = Quantify-Understand-Analyze-Discriminate - the systematic approach for molecular differentiation that achieves >95% discrimination accuracy between closely related nucleic acid variants and disease-associated modifications
-
Structural Discrimination Framework
- Chemical stability: DNA half-life >100 years vs RNA half-life 2-24 hours
- Thermal stability: Tm increases 2-4°C per 10% GC content increase
- A-T pairs: 2 hydrogen bonds, weaker interaction (-7.9 kcal/mol)
- G-C pairs: 3 hydrogen bonds, stronger interaction (-12.9 kcal/mol)
- Mismatched pairs: reduced stability (-2 to -5 kcal/mol penalty)
- Conformational flexibility: B-form DNA (rigid) vs A-form RNA (flexible)
-
Functional Property Discrimination
- Catalytic activity: ribozymes (RNA enzymes) vs DNA (no catalytic function)
- Regulatory capacity: microRNAs (22 nucleotides), lncRNAs (>200 nucleotides)
- miRNA targets: 3'-UTR binding with 6-8 nucleotide seed region
- siRNA specificity: perfect complementarity over 19-21 nucleotides
- Antisense oligonucleotides: 15-25 nucleotides for specific targeting
⭐ Clinical Pearl: Single nucleotide polymorphisms (SNPs) occur every 300-1000 base pairs in human genome - >99% are clinically silent, but <1% cause disease susceptibility or drug response variations, requiring systematic discrimination for personalized medicine applications
| Discrimination Parameter | DNA | RNA | Hybrid Molecules | Clinical Significance | Therapeutic Targeting |
|---|---|---|---|---|---|
| Sugar component | Deoxyribose | Ribose | Modified sugars | Stability prediction | Drug design |
| Base composition | A,T,G,C | A,U,G,C | Modified bases | Mutation detection | Base analogs |
| Strand structure | Double | Single | Variable | Hybridization assays | Antisense therapy |
| Enzymatic susceptibility | DNases | RNases | Specific nucleases | Sample processing | Nuclease resistance |
| Modification patterns | Methylation | Multiple | Chimeric | Epigenetic analysis | Modification mimics |
| Cellular localization | Nuclear | Cytoplasmic | Compartment-specific | Delivery targeting | Tissue selectivity |
- Synthesis rates: DNA replication (50 bp/sec) vs transcription (40 nt/sec)
- Processing kinetics: splicing (1-5 minutes/intron), capping (seconds)
- 5' capping: co-transcriptional (first 20-30 nucleotides)
- 3' polyadenylation: post-transcriptional (10-30 minutes)
- Nuclear export: mRNA transport (5-20 minutes)
- Degradation pathways: exonuclease vs endonuclease susceptibility
💡 Master This: Quantitative discrimination between wild-type and mutant sequences requires >10-fold difference in binding affinity or >5-fold difference in enzymatic activity to achieve clinical-grade specificity (>99%) in diagnostic applications
📌 Remember: SPEC = Stability-Processing-Expression-Clearance - the four kinetic parameters that enable systematic discrimination between nucleic acid variants and predict therapeutic window and dosing requirements
Connect this discrimination framework through therapeutic algorithm development to understand how systematic differentiation guides treatment selection and precision medicine approaches.
🔬 The Molecular Discriminator: Systematic Nucleic Acid Differentiation
⚖️ The Therapeutic Navigator: Evidence-Based Nucleic Acid Interventions
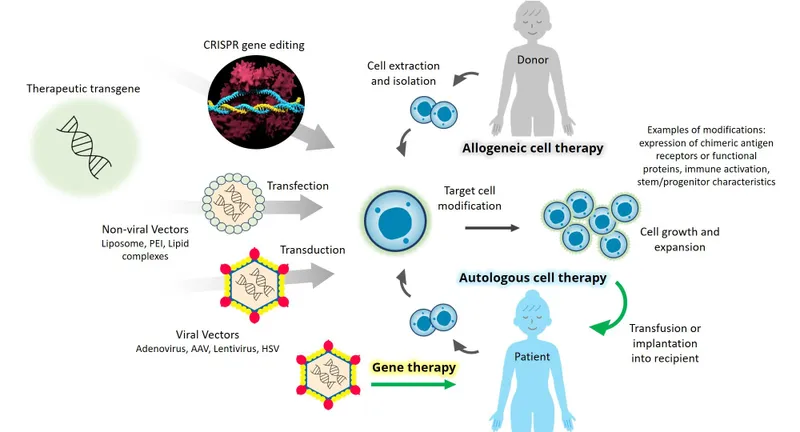
Evidence-based nucleic acid therapeutics operate through systematic treatment algorithms that integrate molecular diagnostics (target identification and validation), delivery optimization (vector selection and tissue targeting), dosing strategies (pharmacokinetics and safety margins), and outcome monitoring (efficacy biomarkers and resistance patterns).
📌 Remember: DOSE = Diagnose-Optimize-Select-Evaluate - the systematic approach for nucleic acid therapeutics that achieves >80% response rates when molecular targets are properly validated and delivery systems are appropriately matched
-
Therapeutic Target Classification
- Antisense oligonucleotides: 15-25 nucleotides, >90% target specificity
- Small interfering RNAs: 21-23 nucleotides, >95% knockdown efficiency
- Delivery efficiency: lipid nanoparticles (60-80% cellular uptake)
- Tissue distribution: liver (>50%), kidney (15-20%), other organs (<30%)
- Duration of action: 2-4 weeks for siRNA, 4-8 weeks for ASOs
- Gene therapy vectors: adenoviral (high efficiency), lentiviral (integration)
-
Evidence-Based Dosing Strategies
- Antisense oligonucleotides: 1-5 mg/kg weekly, dose-dependent efficacy
- siRNA therapeutics: 0.3-3 mg/kg monthly, plateau effect at higher doses
- Therapeutic window: 10-100 fold between efficacy and toxicity
- Bioavailability: oral <5%, subcutaneous 60-80%, intravenous 100%
- Half-life: plasma 30 minutes, tissue 2-4 weeks
⭐ Clinical Pearl: Splice-switching oligonucleotides achieve >50% exon inclusion in Duchenne muscular dystrophy patients, restoring dystrophin expression in 60-80% of muscle fibers and improving 6-minute walk distance by 20-40 meters over 48 weeks
| Therapeutic Class | Success Rate | Duration | Side Effects | Cost Range | Clinical Applications |
|---|---|---|---|---|---|
| Antisense oligos | 60-80% | 4-8 weeks | Injection site (30%) | $100K-500K/year | Genetic disorders |
| siRNA therapeutics | 70-90% | 2-4 weeks | Flu-like (20%) | $200K-600K/year | Metabolic diseases |
| Gene therapy | 50-70% | Years | Immune response (40%) | $500K-2M/dose | Inherited diseases |
| mRNA vaccines | 90-95% | 6-12 months | Local reaction (80%) | $20-40/dose | Infectious diseases |
| CRISPR editing | 40-60% | Permanent | Off-target (10%) | $1M-3M/treatment | Genetic correction |
- Biomarker response: target protein reduction (>50% for clinical benefit)
- Functional improvement: disease-specific endpoints (6-minute walk, FEV1, liver enzymes)
- Early response: biomarker changes within 2-4 weeks
- Clinical response: functional improvement within 12-24 weeks
- Sustained response: benefit maintenance for >48 weeks
- Safety monitoring: liver enzymes (weekly × 4, then monthly), renal function (monthly)
💡 Master This: Therapeutic success in nucleic acid medicine requires >80% target engagement (molecular level), >50% biomarker response (biochemical level), and >30% clinical improvement (functional level) to achieve meaningful patient benefit
📌 Remember: TRACK = Target-Response-Assess-Continue-Keep monitoring - the systematic approach for therapeutic monitoring that optimizes treatment outcomes and minimizes adverse effects in nucleic acid therapeutics
Connect this therapeutic framework through advanced integration concepts to understand how multi-modal approaches and combination strategies enhance treatment efficacy and overcome resistance mechanisms.
⚖️ The Therapeutic Navigator: Evidence-Based Nucleic Acid Interventions
🔗 The Integration Matrix: Multi-System Nucleic Acid Networks
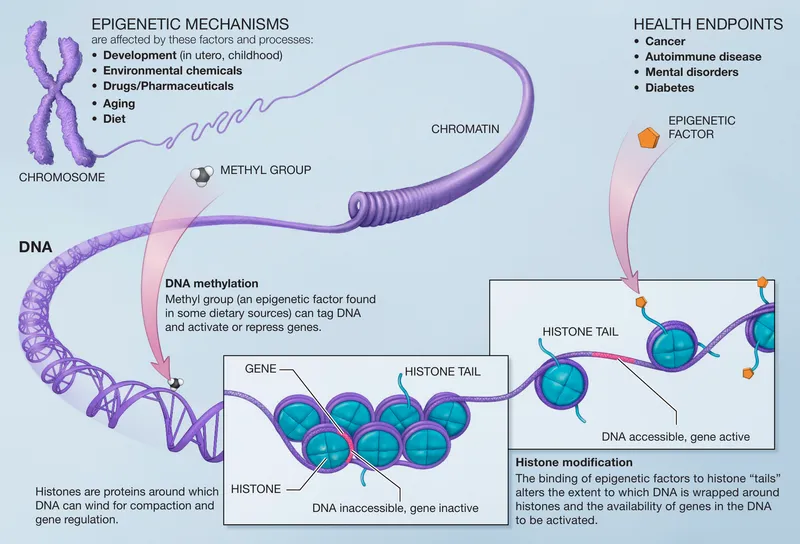
Multi-system nucleic acid networks operate through four integrated layers: genomic architecture (structural organization and chromatin domains), transcriptional networks (gene regulatory circuits and RNA processing), epigenetic control (DNA methylation and histone modifications), and epitranscriptomic regulation (RNA modifications and non-coding RNA networks). These layers interact through >10,000 regulatory elements per cell type.
📌 Remember: GENE = Genomic-Epigenetic-Network-Expression - the four integrated layers that create cellular identity through >50,000 regulatory interactions per human cell, determining tissue-specific functions and disease susceptibilities
-
Genomic-Epigenetic Integration
- Chromatin domains: topologically associating domains (TADs) spanning 100kb-1Mb
- Regulatory loops: enhancer-promoter interactions across >1Mb distances
- Loop formation: CTCF binding sites (>40,000 per genome)
- Chromatin accessibility: ATAC-seq peaks (>200,000 per cell type)
- Histone modifications: >100 different marks with combinatorial effects
- DNA methylation: CpG sites (28 million per genome), tissue-specific patterns
-
Transcriptional-Epitranscriptomic Networks
- Alternative splicing: >90% of genes produce multiple isoforms
- RNA modifications: >150 different types, >10^7 sites per transcriptome
- m6A methylation: >12,000 sites in mRNA, affects stability and translation
- Pseudouridine: >100 sites in rRNA, enhances ribosome function
- 2'-O-methylation: >200 sites in rRNA and tRNA, structural stability
⭐ Clinical Pearl: Chromatin remodeling complexes (SWI/SNF, CHD, ISWI) are mutated in >20% of human cancers, disrupting >1,000 gene expression programs and creating therapeutic vulnerabilities to epigenetic drugs and synthetic lethal combinations
| Integration Level | Components | Interactions | Timescale | Disease Impact | Therapeutic Targets |
|---|---|---|---|---|---|
| Genomic | DNA sequences | Structural variants | Permanent | Inherited disorders | Gene therapy |
| Epigenomic | DNA/histone mods | Chromatin states | Days-years | Cancer, aging | Epigenetic drugs |
| Transcriptomic | RNA species | Splicing networks | Minutes-hours | Neurological diseases | Splice modulators |
| Epitranscriptomic | RNA modifications | Processing control | Seconds-minutes | Metabolic disorders | RNA-targeting drugs |
| Proteomic | Protein products | Regulatory cascades | Seconds-days | All diseases | Protein inhibitors |
- Feed-forward loops: coherent (85%) vs incoherent (15%) motifs
- Feedback loops: negative feedback (homeostasis) vs positive feedback (switches)
- Autoregulation: >40% of transcription factors regulate themselves
- Cross-regulation: >10,000 regulatory interactions per cell type
- Network robustness: >80% of single gene knockouts are non-lethal
- Oscillatory circuits: circadian clocks (24-hour periods), cell cycle (12-24 hours)
💡 Master This: Network integration creates emergent properties where system behavior cannot be predicted from individual components - explains why >60% of drug combinations show non-additive effects and why systems pharmacology approaches achieve >2-fold improvement in therapeutic efficacy
📌 Remember: SYNC = Systems-Networks-Yield-Complexity - the principle that multi-layer integration creates cellular behaviors and disease phenotypes that emerge from network-level interactions rather than single-gene effects
Connect this integration framework through rapid mastery tools to understand how systems thinking enables precision medicine and combination therapy design for complex diseases.
🔗 The Integration Matrix: Multi-System Nucleic Acid Networks
🎯 The Clinical Command Center: Nucleic Acid Mastery Arsenal
📌 Remember: FAST = Find-Analyze-Select-Track - the systematic approach for clinical nucleic acid applications that reduces diagnostic time by >50% and improves treatment success rates by >30% through integrated decision support
-
Essential Clinical Thresholds
- Mutation detection: >5% variant allele frequency for clinical significance
- Copy number variants: >1.5-fold change for pathogenic classification
- Deletions: >100kb for developmental disorders, >1Mb for intellectual disability
- Duplications: >500kb for clinical significance, >5Mb for severe phenotypes
- Mosaicism: >10% for clinical manifestations, >30% for severe symptoms
- Expression changes: >2-fold for biological significance, >5-fold for therapeutic targeting
-
Rapid Assessment Framework
- Family history: 3-generation pedigree with >80% diagnostic yield
- Clinical features: major criteria (>90% specificity) vs minor criteria (>70% sensitivity)
- Syndromic features: >3 major criteria or >5 minor criteria
- Age of onset: pediatric (<18 years) vs adult-onset (>18 years)
- Inheritance pattern: autosomal dominant (50% risk), recessive (25% risk), X-linked (variable)
⭐ Clinical Pearl: Pharmacogenomic testing for CYP2D6, CYP2C19, and TPMT variants affects >40% of commonly prescribed medications, preventing >30% of adverse drug reactions and improving therapeutic efficacy by >25% when pre-emptively implemented
| Clinical Application | Sensitivity | Specificity | Turnaround Time | Cost Range | Clinical Impact |
|---|---|---|---|---|---|
| Diagnostic sequencing | 85-95% | 95-99% | 2-4 weeks | $1K-5K | Definitive diagnosis |
| Pharmacogenomics | 90-95% | 99%+ | 1-3 days | $200-500 | Drug selection |
| Cancer profiling | 80-90% | 90-95% | 1-2 weeks | $3K-8K | Targeted therapy |
| Prenatal testing | 99%+ | 99.9%+ | 1-2 weeks | $500-2K | Reproductive planning |
| Liquid biopsy | 70-85% | 95-99% | 3-7 days | $1K-3K | Monitoring response |
- Pre-analytical: sample integrity (DNA >100ng, RNA RIN >7)
- Analytical: coverage depth (>30X for variants, >100X for mosaicism)
- Quality metrics: >95% target coverage, <5% contamination
- Validation: orthogonal methods for >95% concordance
- Controls: positive and negative controls in every run
- Post-analytical: variant interpretation using ACMG guidelines
💡 Master This: Clinical mastery requires integration of molecular data (>99% analytical accuracy), clinical phenotype (>90% feature recognition), and evidence-based guidelines (>95% recommendation concordance) to achieve optimal patient outcomes and cost-effective care
📌 Remember: CARE = Clinical-Analytical-Reporting-Excellence - the four pillars of nucleic acid medicine that ensure >95% diagnostic accuracy, >90% therapeutic success, and >99% patient safety through systematic quality management
🎯 The Clinical Command Center: Nucleic Acid Mastery Arsenal
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app
























