CRISPR-Cas9: Core Components - Bacterial Defense Force
- CRISPR Array: Bacterial DNA locus with "spacer" (viral memory) and "repeat" sequences.
- Cas Proteins: CRISPR-associated enzymes. Cas9 is a key nuclease (DNA cutter). Cas1/Cas2 acquire spacers.
- RNAs for Targeting:
- crRNA (CRISPR RNA): Contains spacer matching target DNA.
- tracrRNA (trans-activating crRNA): Binds crRNA & Cas9, essential for Cas9 function.
- PAM (Protospacer Adjacent Motif): Short DNA sequence (e.g., 5'-NGG-3') on target DNA, vital for Cas9 activity. Absent in host CRISPR locus.
⭐ PAM sequence is critical for Cas9 to differentiate invader DNA from the bacterial host's CRISPR array.

Mechanism of Genome Editing - Molecular Scalpel
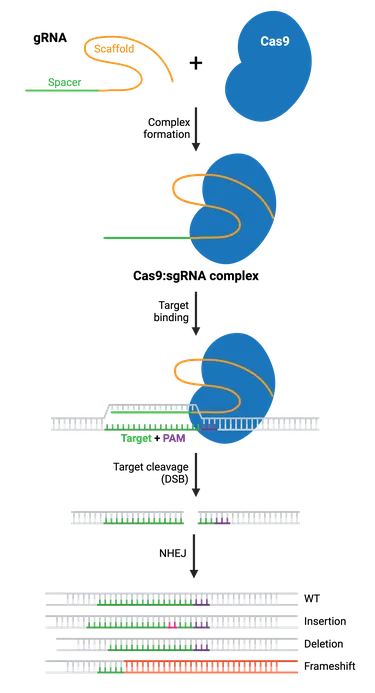
- Key Players:
- Cas9 Protein: An RNA-guided DNA endonuclease. Creates precise DSBs.
- Guide RNA (gRNA): Chimeric RNA; ~20 nt spacer sequence for target DNA binding & scaffold for Cas9 interaction.
- Critical Steps & Outcomes:
- Targeting: gRNA directs Cas9 to DNA. PAM sequence (e.g., 5'-NGG-3') essential for Cas9 activity.
- Cleavage: Cas9 (HNH & RuvC domains) induces DSB.
- Repair Pathways & Results:
- NHEJ: Predominant, error-prone. Leads to indels (gene disruption/knockout).
- HDR: Less frequent, template-dependent. Enables precise edits (gene correction/knock-in).
⭐ Cas9 requires a PAM (Protospacer Adjacent Motif) sequence (e.g., 5'-NGG-3' for Streptococcus pyogenes Cas9) immediately downstream of the gRNA target sequence for DNA cleavage.
Therapeutic & Research Applications - Fixing Faulty Genes
- Gene Therapy:
- Corrects monogenic disorders: Sickle Cell Disease (SCD), β-thalassemia, Cystic Fibrosis (CF), Duchenne Muscular Dystrophy (DMD).
- Utilizes ex-vivo (e.g., hematopoietic stem cells - HSCs) & in-vivo editing strategies.
- Oncology:
- CAR-T cell engineering for potent cancer immunotherapy.
- Targeting oncogenes or restoring tumor suppressor gene functions.
- Infectious Diseases:
- Antiviral strategies (e.g., HIV, HBV); developing novel antimicrobials against resistant pathogens.
- Research & Diagnostics:
- Creating precise disease models (cellular, animal) for study.
- Functional genomics: studying gene functions and biological pathways.
- Drug discovery and validating new therapeutic targets.
- Developing rapid and sensitive molecular diagnostic tools.
⭐ Key clinical trials involve ex-vivo CRISPR editing of hematopoietic stem cells (HSCs) for β-thalassemia and Sickle Cell Disease to restore functional hemoglobin production.OKA
Challenges & Ethical Landscape - Navigating the Maze
- Technical Challenges: (📌 O M D I)
- Off-target effects: unintended genome edits.
- Mosaicism: mixed cell editing outcomes.
- Delivery: efficient in vivo vector systems.
- Immunogenicity: against Cas/vector.
- Ethical, Legal, Social Implications (ELSI):
- Germline (heritable) vs. Somatic (therapeutic) editing.
- Enhancement vs. Therapy debate.
- Access, equity, and justice.
- Informed consent for novel therapies.
⭐ Key challenge: Minimizing off-target mutations for clinical safety.
Advanced CRISPR Technologies - Beyond Basic Editing
- Alternative Cas enzymes:
- Cas12a (Cpf1): Staggered cuts, T-rich PAM.
- Cas13: RNA targeting (editing, knockdown).
- Base Editing (BE):
- Cytidine Base Editors (CBEs): C•G → T•A.
- Adenine Base Editors (ABEs): A•T → G•C.
- Precise, no Double-Strand Breaks (DSBs).
- Prime Editing (PE): Versatile "search-and-replace" editing; no DSBs or donor DNA.
- dCas9 (dead Cas9): Nuclease-deficient; for transcriptional regulation (CRISPRi/a), imaging.
⭐ Prime editing allows all 12 possible base-to-base conversions, plus small insertions and deletions, without requiring double-strand breaks or donor DNA templates, offering high precision and versatility.
High‑Yield Points - ⚡ Biggest Takeaways
CRISPR refers to Clustered Regularly Interspaced Short Palindromic Repeats. Cas9, an endonuclease, is guided by gRNA (crRNA + tracrRNA complex) to target DNA. gRNA directs Cas9 to create specific double-strand breaks (DSBs). Cellular repair of DSBs occurs via NHEJ (error-prone) or HDR (precise, uses template). PAM (Protospacer Adjacent Motif) sequence (e.g., NGG for SpCas9) is vital for Cas9 binding and cleavage. Key applications: gene editing, knockouts, insertions, and transcriptional regulation.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

