Ribozymes - RNA's Cutting Edge
- RNA molecules with catalytic activity; "catalytic RNA".
- Discovered by Altman & Cech (Nobel Prize). Challenged protein-only enzyme dogma.
- Key Examples:
- Self-splicing introns (Group I, II).
- RNase P (tRNA processing).
- Ribosomal RNA (rRNA): forms peptide bonds.
- Supports RNA World hypothesis (RNA as early catalyst & genetic material).
- Often use divalent metal ions (e.g., $Mg^{2+}$) as cofactors.
⭐ Peptidyl transferase center of the ribosome is rRNA, not protein, highlighting RNA's catalytic role in protein synthesis.
Catalytic Action - How RNA Slices
- Core Principle: RNA folds into complex 3D structures, forming active sites.
- Relies on specific nucleotide sequences.
- Utilizes 2'-OH groups, bases, and phosphate backbone for catalysis.
- Metal Ion Cofactors: Crucial, especially $Mg^{2+}$.
- Structural role: Stabilize RNA tertiary structure.
- Catalytic role: Act as Lewis acids, activate nucleophiles (e.g., $H_2O$), stabilize transition states & leaving groups.
- Mechanism of "Slicing" (Phosphodiester Bond Hydrolysis/Transesterification):
- Nucleophilic Attack: Often by an activated hydroxyl group (RNA's 2'-OH, 3'-OH, or an external $H_2O$).
- Transition State Stabilization: By metal ions and/or RNA functional groups.
- Leaving Group Departure: Facilitated to complete bond cleavage.
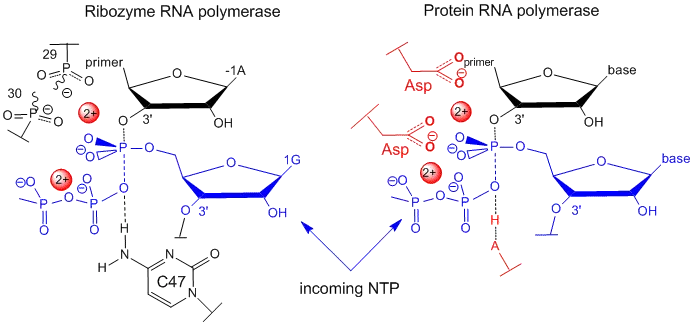
- Common: Two-metal-ion mechanism often employed (e.g., Group I introns, RNase H-like).
⭐ The catalytic RNA component of RNase P is essential for the maturation of precursor tRNA molecules by cleaving the 5' leader sequence, a classic example of RNA slicing.
Ribozyme Roster - Key RNA Players
- Self-Splicing Introns: RNA molecules that excise themselves from a larger RNA transcript.
- Group I: External guanosine ($G$) cofactor dependent. Found in bacteria, lower eukaryotes, plants (rRNA, mRNA, tRNA).
- Group II: Internal adenosine ($A$) initiates lariat formation. Mechanistically similar to spliceosomal introns. Found in organellar genes, prokaryotes.
- RNase P: Ribonucleoprotein; RNA subunit is catalytic. Processes 5' end of pre-tRNAs.
⭐ The RNA component of bacterial RNase P can catalyze tRNA maturation in vitro without its protein cofactor.
- Ribosome (Peptidyl Transferase Center): rRNA in the large subunit catalyzes peptide bond formation during protein synthesis.
- Small Viral/Viroid Ribozymes: Site-specific RNA cleavage, crucial for replication.
- Hammerhead Ribozyme: Viroids, satellite RNAs.
- Hairpin Ribozyme: Satellite RNAs.
- Hepatitis Delta Virus (HDV) Ribozyme: Self-cleaving in HDV genome.
- Spliceosomal snRNAs (e.g., U2, U6): Core components of the spliceosome, implicated in catalytic activity during pre-mRNA splicing.
 , RNase P, ribosomal PTC, hammerhead ribozyme)
, RNase P, ribosomal PTC, hammerhead ribozyme)
RNA Rx - Ribozymes in Medicine
- Therapeutic Potential:
- Gene silencing: Engineered to cleave specific viral or oncogenic mRNAs.
- Common types: Hammerhead, hairpin ribozymes.
- Targets: HIV, HCV, cancer (HER2), macular degeneration (Angiozyme for VEGF mRNA).
- Clinical Hurdles:
- In vivo delivery, nuclease stability, off-target effects.
- Other Applications:
- Diagnostic biosensors (detecting nucleic acids/metabolites).
- Biotechnology tools for RNA research.
⭐ Angiozyme, an engineered hairpin ribozyme, targets VEGF mRNA and was tested in clinical trials for age-related macular degeneration.
High‑Yield Points - ⚡ Biggest Takeaways
- Ribozymes: RNA molecules with catalytic activity; not exclusively proteins.
- Examples: RNase P (tRNA maturation), self-splicing introns, ribosomal RNA (peptidyl transferase).
- Small ribozymes: Hammerhead, hairpin, HDV; vital for viral replication via self-cleavage.
- Often require divalent metal ions (e.g., Mg²⁺) as cofactors for catalysis.
- Support RNA world hypothesis: RNA as early biocatalyst and genetic material.
- Therapeutic potential: gene therapy targeting specific RNAs.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

