Enzyme Essentials - Name Game Gurus
- Enzymes: Biological catalysts, typically proteins, accelerating biochemical reactions without being consumed.
- Active Site: Specific 3D pocket on the enzyme where the substrate binds and catalysis occurs.
- Substrate (S): The reactant molecule(s) upon which an enzyme acts.
- Enzyme Components:
- Apoenzyme: The protein part of an enzyme; inactive on its own.
- Cofactor: Non-protein chemical compound (ion or molecule) required for an enzyme's activity.
- Coenzyme: Organic, loosely bound cofactor (e.g., vitamins like NAD+, FAD+).
- Prosthetic Group: Organic or inorganic cofactor, tightly bound to the apoenzyme (e.g., heme, FMN).
- Holoenzyme: The complete, catalytically active enzyme, consisting of an apoenzyme and its cofactor. 📌 Holo = Whole (active).
- Nomenclature:
- Trivial names: Common, often historical (e.g., Pepsin, Trypsin).
- Systematic names: Based on the substrate and type of reaction catalyzed, ending with the suffix "-ase" (e.g., Lactate dehydrogenase).
⭐ Ribozymes are catalytic RNAs, an exception to the general rule that enzymes are proteins.
Enzyme Classes 1 & 2 - Redox & Group Shuttlers
📌 Mnemonic for 6 enzyme classes: OTHLIL (Oscar The Hairy LILy)
| Class | EC No. | Reaction Type & Equation | Examples & Specifics | Coenzymes / Groups Transferred |
|---|---|---|---|---|
| Oxidoreductases | EC 1 | Catalyze redox reactions (electron transfer). $S_{red} + A_{ox} \rightleftharpoons S_{ox} + A_{red}## Enzyme Classes 1 & 2 - Redox & Group Shuttlers |
📌 Mnemonic for 6 enzyme classes: OTHLIL (Oscar The Hairy LILy)
| Dehydrogenases (remove H), oxidases (use $O_2$ as acceptor), peroxidases. | NAD+/NADP+, FAD/FMN as coenzymes. | | Transferases | EC 2 | Transfer functional groups (e.g., methyl, amino, phosphate). $G-X + A \rightleftharpoons G-A + X## Enzyme Classes 1 & 2 - Redox & Group Shuttlers
📌 Mnemonic for 6 enzyme classes: OTHLIL (Oscar The Hairy LILy)
| Kinases (transfer $PO_4^{3-}$), transaminases (e.g., ALT, AST; transfer amino group). | C, N, P-containing groups. |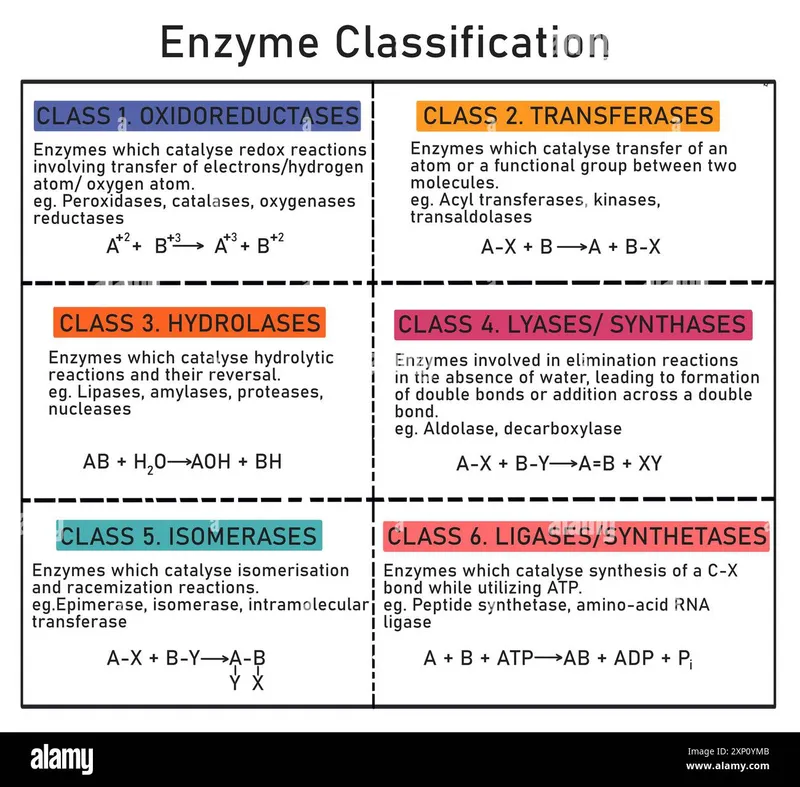
⭐ Alanine aminotransferase (ALT) is a more specific indicator of liver damage than Aspartate aminotransferase (AST).
Enzyme Classes 3 & 4 - Water Breakers & Bond Splitters
| Class (EC) | Name | Reaction Type & Mechanism | Examples |
|---|---|---|---|
| EC 3 | Hydrolases | Hydrolysis: Cleave bonds by adding water. $A-B + H_2O \rightarrow A-OH + B-H## Enzyme Classes 3 & 4 - Water Breakers & Bond Splitters |
| Peptidases, lipases, nucleases, glycosidases | | EC 4 | Lyases | Non-hydrolytic/oxidative cleavage of C-C, C-O, C-N bonds; forms double bonds/rings. $X-A-B-Y \rightarrow A=B + X-Y## Enzyme Classes 3 & 4 - Water Breakers & Bond Splitters
| Decarboxylases, aldolases, dehydratases, synthases* |* 📌 OTHLIL: Hydrolases, Lyases.
- Synthases (Lyases) cleave bonds; Synthetases (Ligases, EC 6) join molecules (ATP-dependent).
⭐ Many digestive enzymes (e.g., pepsin, amylase, lipase) are hydrolases, crucial for nutrient breakdown.
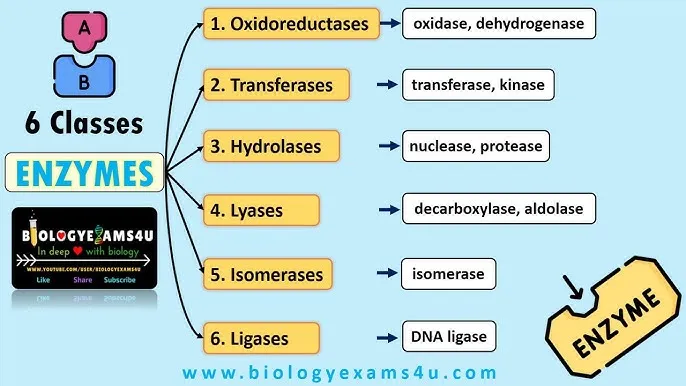
Enzyme Classes 5 & 6 & EC Codes - Shifters, Joiners & ID Tags
📌 OTHLIL: Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases.
EC 5: Isomerases ("Shifters")
- Intramolecular rearrangements (geometry/structure change). Reaction: $A-B-C \rightarrow A-C-B$.
- E.g., L-alanine $\leftrightarrow$ D-alanine. Types: Epimerases, racemases, mutases.
EC 6: Ligases ("Joiners")
- Join two molecules; form new C-O, C-S, C-N, C-C bonds; ATP-dependent. Reaction: $A + B + ATP \rightarrow A-B + ADP + P_i$.
- Types: Synthetases, carboxylases. E.g., DNA ligase.
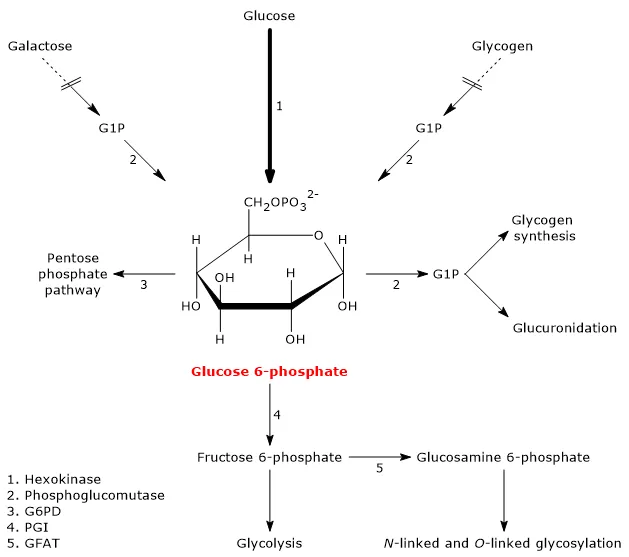
⭐ DNA Ligase (EC 6), essential for DNA replication and repair by joining DNA fragments, requires ATP.
EC Number System (ID Tags): EC W.X.Y.Z
- W: Main class (enzyme's broad category, 1-7).
- X: Subclass (type of bond/group acted upon).
- Y: Sub-subclass (further details like cofactor/acceptor).
- Z: Serial number (specific enzyme's unique ID).
High‑Yield Points - ⚡ Biggest Takeaways
- Six major enzyme classes (EC 1-6) are defined by the Enzyme Commission (IUBMB): Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases (OTHLIL).
- Each enzyme has a unique four-digit EC number (Class.Subclass.Sub-subclass.Specific enzyme).
- Oxidoreductases (EC 1) catalyze redox reactions; Transferases (EC 2) move functional groups.
- Hydrolases (EC 3) cleave bonds using water; Lyases (EC 4) cleave bonds non-hydrolytically, often forming double bonds.
- Isomerases (EC 5) rearrange atoms within a molecule.
- Ligases (EC 6) join two molecules, typically using ATP hydrolysis.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

