Overview & Tools - Gene Splicing Starter Kit
- Recombinant DNA Technology (RDT): Techniques for manipulating DNA by joining DNA segments from different sources in vitro to create a recombinant DNA molecule, which is then introduced into a host organism.
- Essential Tools ("Gene Splicing Starter Kit"):
- Enzymes - The Molecular Toolkit:
- Restriction Endonucleases: "Molecular scissors"; recognize & cleave specific DNA sequences (palindromic sites). E.g., EcoRI, HindIII. Create sticky or blunt ends.
- DNA Ligase: "Molecular glue"; forms phosphodiester bonds to join DNA fragments.
- DNA Polymerase (e.g., Taq polymerase for PCR): DNA synthesis.
- Reverse Transcriptase: mRNA → cDNA (complementary DNA).
- Vectors: Cloning vehicles (e.g., plasmids, bacteriophages, cosmids, BACs, YACs) to carry gene of interest into host. Must have origin of replication (ori), selectable marker, cloning sites.
- Host Organism: For replication & expression of recombinant DNA (e.g., E. coli, yeast).
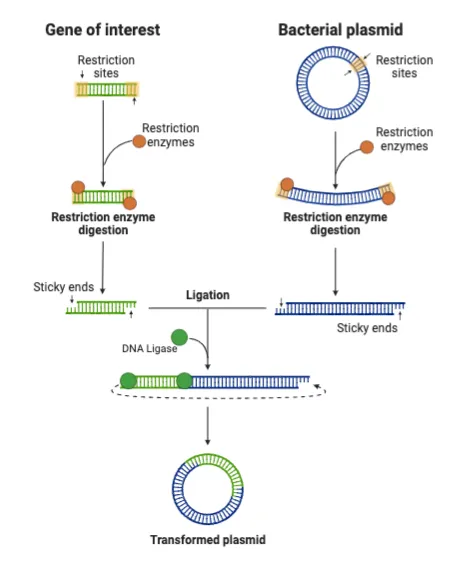
- Enzymes - The Molecular Toolkit:
⭐ Type II restriction endonucleases are most commonly used in RDT as they cleave DNA at specific sites within or very near their recognition sequence, requiring no ATP for their activity (unlike Type I & III).
Cloning Vectors - DNA Delivery Vans
- Role: Carry & replicate foreign DNA in host cells.
- Key Features:
- ori: For self-replication.
- Selectable Marker: E.g., antibiotic resistance (like $Amp^R$ for ampicillin, $Tet^R$ for tetracycline); identifies transformants.
- Multiple Cloning Site (MCS): For inserting DNA.
- Small Size: Efficient transfer.
- Types & Capacity:
- Plasmids (pBR322, pUC): < 15 kb.
- Bacteriophages (λ, M13): < 25 kb.
- Cosmids: < 45 kb.
- BACs (Bacterial Artificial Chromosomes): 100-300 kb.
- YACs (Yeast Artificial Chromosomes): 200-2000 kb.
- Expression Vectors: For protein synthesis (need promoter, RBS).

⭐ pBR322 utilizes its ampicillin and tetracycline resistance genes for selection; insertional inactivation in one of these helps screen for recombinants.
Core RDT Procedures - Genetic Engineering Toolkit
Essential tools enabling gene manipulation:
- Enzymes (Molecular Machinery):
- Restriction Endonucleases: "Molecular scissors" cleaving DNA at specific palindromic sites (e.g., EcoRI at GAATTC). Type II pivotal.
- DNA Ligase: "Molecular glue" joining DNA fragments (phosphodiester bonds).
- DNA Polymerases: Synthesize DNA (e.g., Taq polymerase in PCR).
- Reverse Transcriptase: mRNA → cDNA.
- Vectors (Cloning Vehicles): Carry foreign DNA into a host.
- Types: Plasmids, Bacteriophages, Cosmids, BACs, YACs.
- Features: Origin of replication (ori), selectable marker, cloning site (MCS).
- Host Organism: Replicates/expresses recombinant DNA (e.g., E. coli).

⭐ Restriction enzymes like EcoRI recognize specific palindromic sequences (e.g., 5'-GAATTC-3'), which read the same forwards on one strand and backwards on the complementary strand.
Applications & Libraries - RDT's Greatest Hits
- Therapeutic Proteins: Insulin (diabetes), erythropoietin (anemia), growth hormone (dwarfism), interferons (antiviral, anticancer), clotting factors (hemophilia).
- Vaccines: Hepatitis B vaccine (recombinant HBsAg), HPV vaccine.
- Diagnostics: DNA probes for genetic disorders (e.g., sickle cell anemia, cystic fibrosis), infectious diseases (e.g., HIV, TB).
- Gene Therapy: Introducing functional genes to treat genetic diseases.
- Transgenic Organisms:
- Plants: Pest resistance (Bt cotton), herbicide tolerance, improved nutritional value (Golden Rice).
- Animals: Disease models, enhanced production (e.g., milk).
- DNA Libraries: Collections of cloned DNA fragments.
- Genomic Library: Represents entire genome.
- cDNA Library: Represents only expressed genes (mRNA transcripts); no introns.
⭐ cDNA libraries are preferred for expressing eukaryotic genes in prokaryotes because prokaryotes cannot splice introns.

- Pharmacogenomics: Tailoring drug therapy based on individual genetic makeup.
- Forensic Science: DNA fingerprinting (VNTRs, STRs).
High‑Yield Points - ⚡ Biggest Takeaways
- Restriction enzymes act as 'molecular scissors', recognizing specific palindromic DNA sequences.
- Cloning vectors like plasmids (e.g., pBR322) and bacteriophages carry foreign DNA.
- DNA ligase is the 'molecular glue' that joins DNA fragments.
- PCR enables rapid DNA amplification using Taq polymerase and specific primers.
- Blotting techniques: Southern (DNA), Northern (RNA), Western (protein) for detection.
- cDNA libraries, derived from mRNA, represent actively transcribed genes.
- Applications include recombinant insulin, vaccines, gene therapy, and disease diagnosis.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

