Proteomics Primer - Protein Power Play
Large-scale study of proteins: their expression levels, structures, functions, post-translational modifications (PTMs), and interactions. Aims to characterize the entire proteome of a cell or organism under specific conditions.
- Core Workflow:
- Key Techniques:
- Separation:
- 2D-PAGE: By isoelectric point (pI, 1st D) & Molecular Weight (MW, 2nd D). Visualizes expression changes.
- Liquid Chromatography (LC): Often coupled with MS (LC-MS).
- Identification & Quantification (ID/Quant):
- Mass Spectrometry (MS): Core technology. Measures mass-to-charge ratio ($m/z$) of ionized peptides.
- MALDI-TOF: Good for simpler mixtures, Peptide Mass Fingerprinting (PMF).
- ESI-MS/MS: For complex mixtures, peptide sequencing (shotgun proteomics).
- Quantitative methods: SILAC, iTRAQ, Label-Free Quantification (LFQ).
- Mass Spectrometry (MS): Core technology. Measures mass-to-charge ratio ($m/z$) of ionized peptides.
- Interaction Analysis: Yeast-2-Hybrid (Y2H), Co-Immunoprecipitation (Co-IP) followed by MS.
- Separation:
- Applications:
- Biomarker discovery (disease diagnosis, prognosis, therapy response).
- Drug target identification & validation.
- Understanding disease pathogenesis (e.g., cancer, neurodegenerative disorders).
- Analysis of PTMs (e.g., phosphorylation, glycosylation).
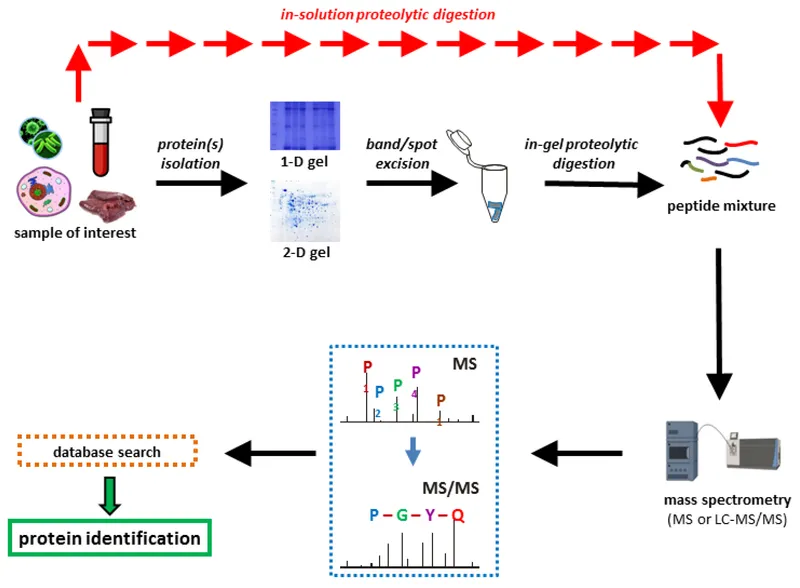
⭐ Shotgun proteomics (LC-MS/MS) identifies thousands of proteins from complex samples without prior 2D-PAGE, revolutionizing biomarker discovery and systems biology research.
Metabolomics Mania - Small Molecule Secrets
Metabolomics is the systematic study of unique chemical fingerprints (metabolites, < 1.5 kDa) specific to cellular processes. It offers a direct snapshot of physiological or pathological states.
- Goal: Identify & quantify the complete metabolome.
- Significance:
- Reflects phenotype directly.
- Captures dynamic cellular responses.
- Key Analytical Platforms:
- Mass Spectrometry (MS): Coupled with Gas/Liquid Chromatography (GC-MS, LC-MS). High sensitivity, broad coverage. Ideal for complex mixtures.
- Nuclear Magnetic Resonance (NMR): Non-destructive, quantitative, good for structural elucidation. Less sensitive than MS.
- Common Samples: Plasma, urine, serum, CSF, tissue biopsies.
- Approaches:
- Untargeted: Global metabolic profiling; hypothesis-generating.
- Targeted: Measures specific, known metabolites; hypothesis-driven.
⭐ Phenylketonuria (PKU), an Inborn Error of Metabolism, is diagnosed by detecting elevated phenylalanine levels using tandem mass spectrometry (MS/MS) in newborn screening - a prime example of targeted metabolomics.
Omics Applications & Synergy - Clinical Clues Unlocked
- Proteomics: Decoding Protein Functions & Interactions
- Biomarker Discovery: Identifying proteins (e.g., Troponins for MI) for early diagnosis, prognosis, and monitoring therapeutic response.
- Drug Development: Crucial for novel drug target identification, validation, and understanding drug mechanisms.
- Personalized Medicine: Tailoring treatments by analyzing individual patient proteomes, predicting drug efficacy.
- Metabolomics: Profiling Small Molecule Metabolites
- Disease Diagnosis & Staging: Revealing metabolic signatures for conditions like diabetes, cancer, and inborn errors of metabolism.
- Pharmacometabolomics: Predicting individual drug responses and adverse effects, guiding personalized drug therapy.
- Nutritional & Environmental Health: Assessing dietary impacts and exposure to toxins.
- Synergy in Systems Medicine:
- Integrated Multi-Omics: Combining proteomics and metabolomics with genomics/transcriptomics offers a comprehensive view of cellular states.
- Pathway Perturbation Analysis: Elucidating disease mechanisms by mapping changes in protein and metabolite networks.
- Clinical Impact: Accelerates development of precise diagnostics, targeted therapies, and preventative strategies.

⭐ Metabolomic profiling of urine can detect early signs of diabetic nephropathy by identifying specific biomarkers like altered amino acids or sugars.
High‑Yield Points - ⚡ Biggest Takeaways
- Proteomics analyzes the entire protein set (proteome); key tools include 2D-PAGE and Mass Spectrometry (MS).
- MALDI-TOF and ESI are crucial MS ionization techniques for protein identification.
- Metabolomics studies all small molecules (metabolites) within a biological system.
- NMR spectroscopy and MS-based methods (GC-MS, LC-MS) are central to metabolomics.
- Both fields are vital for biomarker discovery and understanding disease mechanisms.
- Systems biology integrates these 'omics' data for a holistic view of cellular processes.
- Protein microarrays and yeast two-hybrid systems are used to study protein interactions in proteomics.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

