Epigenetics Overview - The Cancer Conductor
Heritable, reversible changes in gene expression without altering the DNA sequence. These modifications can silence tumor suppressor genes or activate oncogenes, acting as a conductor for carcinogenesis.
- Key Mechanisms:
- DNA Methylation: Hypermethylation of CpG islands in promoter regions → silences tumor suppressor genes (e.g., CDKN2A).
- Histone Modification: Altered acetylation/deacetylation patterns change chromatin accessibility and gene expression.
⭐ Epigenetic alterations are reversible, forming a basis for novel cancer therapies (e.g., DNMT and HDAC inhibitors).
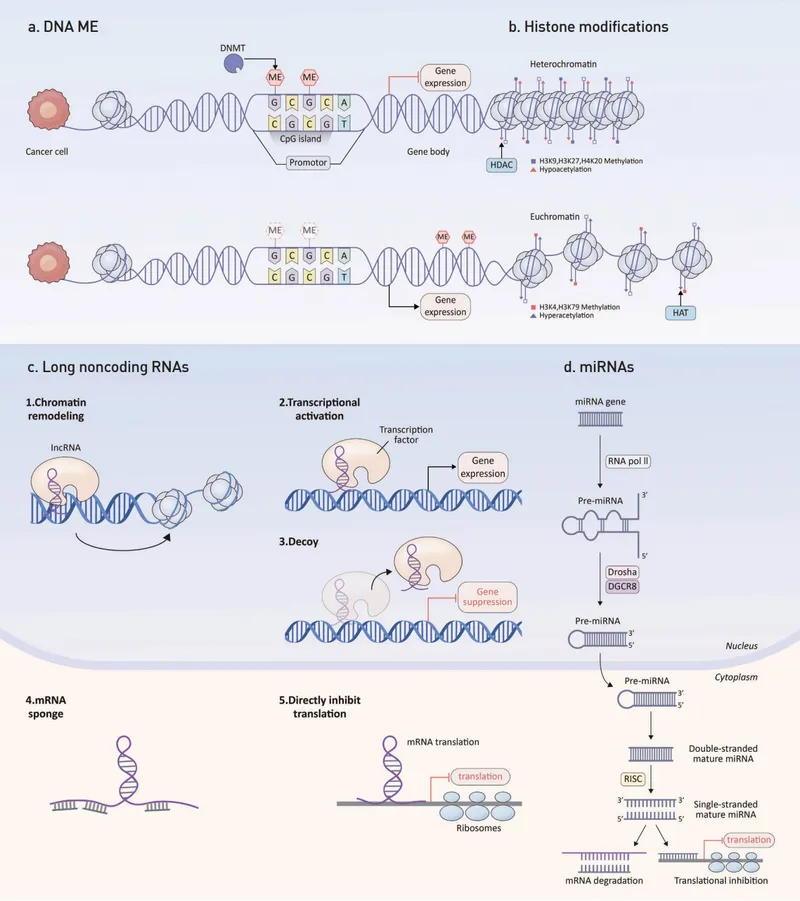
DNA Methylation - The Master Switch
- Mechanism: Covalent addition of a methyl group ($CH_3$) to cytosine, primarily at CpG dinucleotides.
- Enzyme: Catalyzed by DNA Methyltransferases (DNMTs).
- Function: Acts as a master switch for gene expression.
- ↑ Methylation (Hypermethylation) → condensed chromatin → gene silencing.
- ↓ Methylation (Hypomethylation) → relaxed chromatin → gene activation.
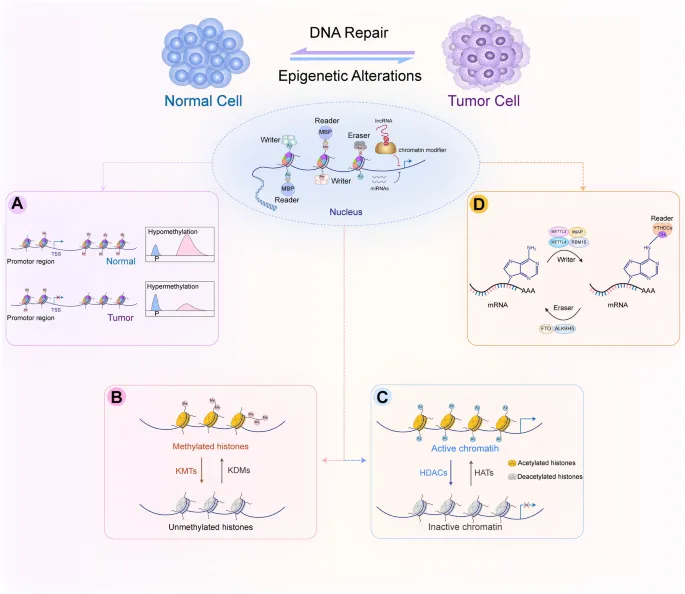
- Role in Cancer:
- Hypermethylation: Silences tumor suppressor genes (e.g., p16, BRCA1).
- Hypomethylation: Activates oncogenes and promotes genomic instability.
⭐ High-Yield: Aberrant hypermethylation of CpG islands in the promoter regions of tumor suppressor genes is a hallmark of many cancers, leading to their transcriptional inactivation.
Histone Modification - Unspooling Trouble
-
Core Concept: Chemical changes to histone proteins alter chromatin structure & gene accessibility. Tightly wound heterochromatin silences genes; loosely packed euchromatin allows transcription.
-
Key Modifications:
- Acetylation: Histone Acetyltransferases (HATs) add acetyl groups → relaxed chromatin (euchromatin) → ↑ gene expression. Histone Deacetylases (HDACs) reverse this.
- Methylation: Histone Methyltransferases (HMTs) add methyl groups. Effect varies: can activate or repress genes depending on the site.
-
Role in Cancer:
- Mutations in genes for HATs/HDACs or HMTs can silence tumor suppressor genes (e.g., p53) or activate oncogenes.
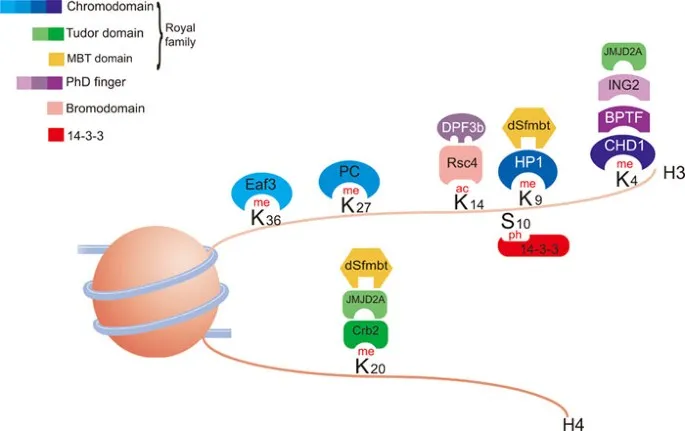
⭐ Exam Favorite: Histone Deacetylase (HDAC) inhibitors (e.g., Vorinostat, Romidepsin) are a class of anti-cancer drugs. They work by promoting histone acetylation, leading to re-expression of silenced tumor suppressor genes.
Epigenetic Drugs - Hacking the Code
Goal: Reactivate tumor suppressor genes silenced by epigenetic modifications.
- DNMT Inhibitors: Azacitidine, Decitabine. Hypomethylating agents that incorporate into DNA, trapping DNMTs and leading to ↓ methylation.
- HDAC Inhibitors: Vorinostat, Romidepsin. Prevent removal of acetyl groups from histones, leading to ↑ acetylation, open chromatin, and gene expression.
⭐ Azacitidine and Decitabine are workhorses for myelodysplastic syndromes (MDS), often improving blood counts and delaying progression to acute myeloid leukemia (AML).
- DNA hypermethylation at CpG islands in promoter regions silences tumor suppressor genes (e.g., MLH1, BRCA1).
- Widespread genomic hypomethylation can lead to chromosomal instability and activation of proto-oncogenes.
- Histone modifications, like deacetylation, can condense chromatin and inhibit transcription of key regulatory genes.
- MicroRNAs (miRNAs) can function as oncogenes or tumor suppressors by regulating mRNA translation.
- Unlike genetic mutations, epigenetic alterations are potentially reversible, offering novel therapeutic targets (e.g., DNMT and HDAC inhibitors).
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

