Replication Essentials - The Ground Rules
- Semiconservative: Each new DNA molecule contains one parent (old) strand and one daughter (new) strand.
- Origin of Replication (ORI):
- Starting point; typically AT-rich (fewer H-bonds).
- Prokaryotes have a single ORI; Eukaryotes have multiple ORIs.
- Bidirectional: Replication proceeds in both directions from the ORI, forming two replication forks.
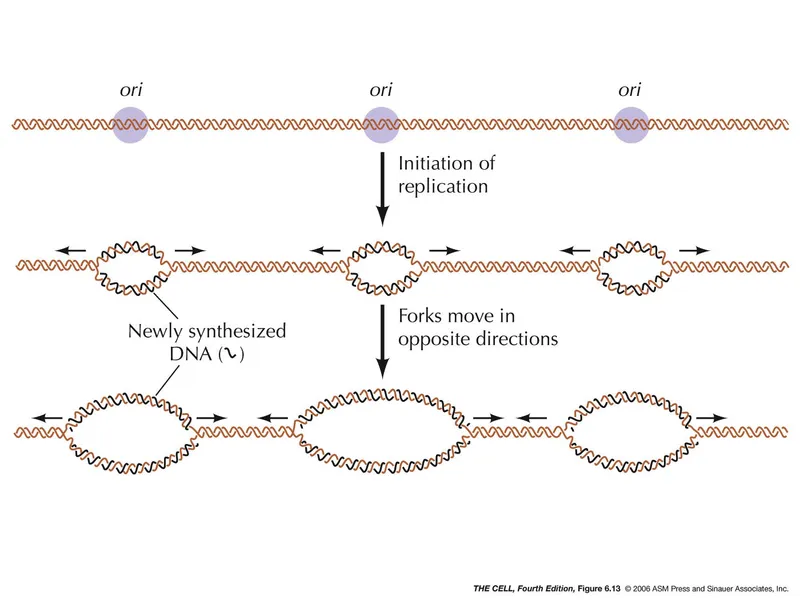
- 5' → 3' Synthesis: DNA polymerase adds nucleotides ONLY to the free 3'-hydroxyl end of a growing strand.
⭐ The 5' → 3' directionality of DNA polymerase is absolute and dictates the existence of a continuous leading strand and a discontinuous lagging strand (Okazaki fragments).
The Replication Fork - Unzipping Action
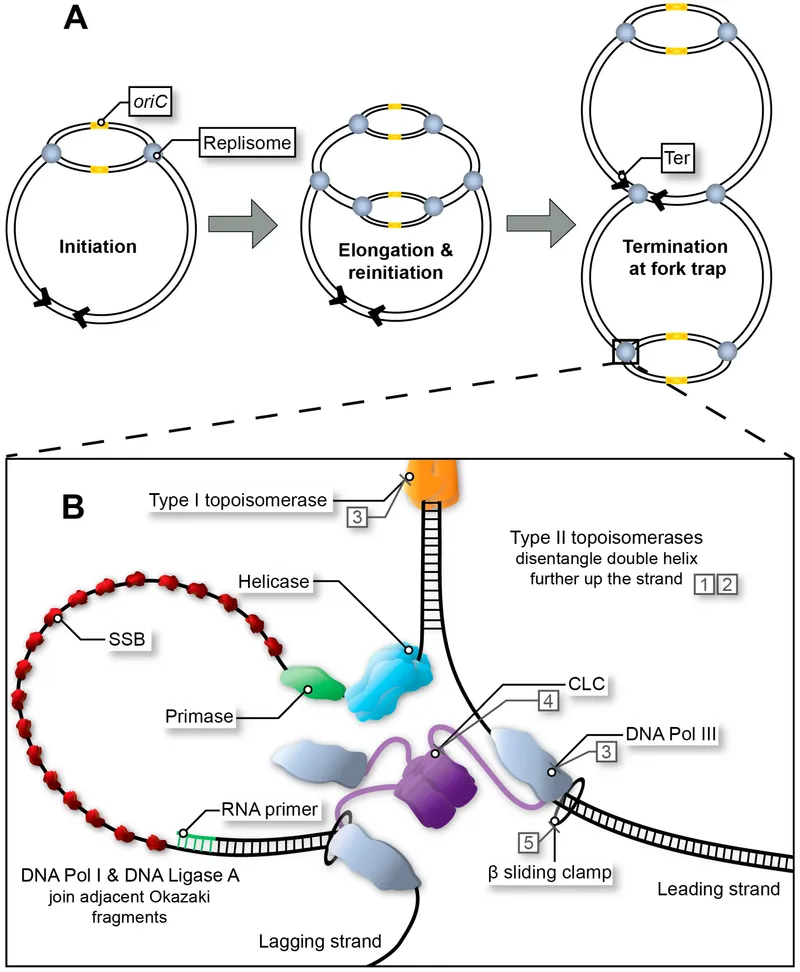
- Helicase: Unzips the DNA double helix at the replication fork, breaking hydrogen bonds in an ATP-dependent process.
- Single-Strand Binding Proteins (SSBPs): Bind to the separated strands to keep them from re-annealing and protect them from nuclease degradation.
- DNA Topoisomerases: Relieve the torsional strain and negative supercoils that build up ahead of the fork.
- Type I: Creates single-strand nicks to relax supercoils.
- Type II (DNA Gyrase in E. coli): Induces double-strand breaks to manage supercoils.
⭐ Fluoroquinolones inhibit prokaryotic Type II topoisomerase (DNA gyrase). Etoposide/teniposide inhibit human Type II topoisomerase.
Prokaryote vs Eukaryote - Tale of Two Cells
- Origin of Replication: Single in prokaryotes vs. multiple in eukaryotes, allowing for faster overall replication despite a slower polymerase speed.
- DNA Polymerases:
- Prokaryotes: Pol I, II, III. 📌 Pol III is the primary replicative enzyme.
- Eukaryotes: Complex (α, δ, ε).
- Telomerase: Absent in prokaryotes (circular DNA); present in eukaryotes to maintain telomere length on linear chromosomes.
- Okazaki Fragments: Longer in prokaryotes; shorter in eukaryotes.
⭐ Eukaryotic DNA Pol α has primase activity and initiates synthesis. Pol δ synthesizes the lagging strand, and Pol ε synthesizes the leading strand.
Telomeres & Drugs - Ends and Blockers
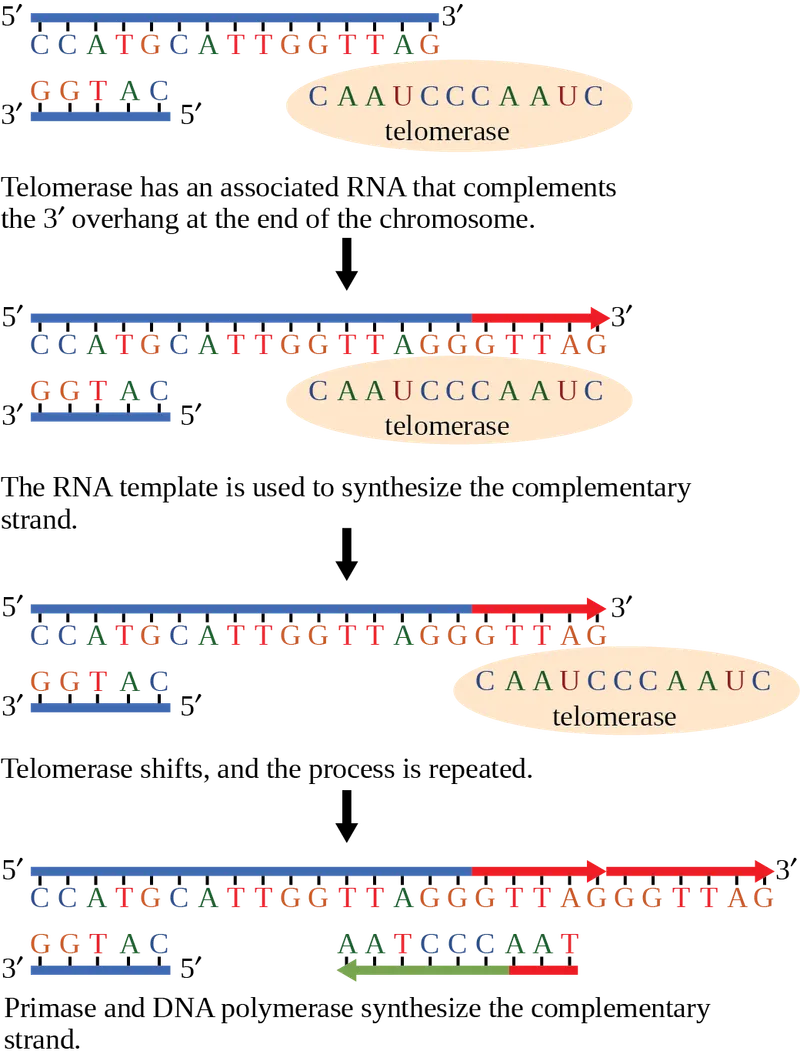
- Telomeres: Repetitive DNA sequences (TTAGGG) at chromosome ends protecting against degradation. They shorten with each cell division, contributing to aging.
- Telomerase: A reverse transcriptase that lengthens telomeres.
- High activity in stem, germ, and cancer cells.
- Uses an internal RNA template to add repeats.
- Replication Inhibitors:
- Hydroxyurea: Blocks ribonucleotide reductase, inhibiting dNTP synthesis.
- 5-FU, Methotrexate: Inhibit thymidylate synthesis.
⭐ Telomerase reactivation is a hallmark of ~90% of human cancers, allowing them to evade senescence and achieve cellular "immortality."
High‑Yield Points - ⚡ Biggest Takeaways
- DNA replication is semiconservative and always proceeds in the 5' to 3' direction.
- Helicase unwinds the DNA helix, while topoisomerases (e.g., DNA gyrase) relieve supercoiling.
- Primase synthesizes an RNA primer to initiate DNA synthesis.
- DNA Polymerase III is the primary enzyme for synthesizing new DNA in prokaryotes.
- The lagging strand is synthesized discontinuously, creating Okazaki fragments.
- DNA Polymerase I removes RNA primers, and DNA ligase joins the fragments.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

